| Excerpt |
|---|
This collection contains images from 422 non-small cell lung cancer (NSCLC) patients. For these patients pretreatment CT scans, manual delineation by a radiation oncologist of the 3D volume of the gross tumor volume and clinical outcome data are available. This dataset refers to the Lung1 dataset of the study published in Nature Communications. |
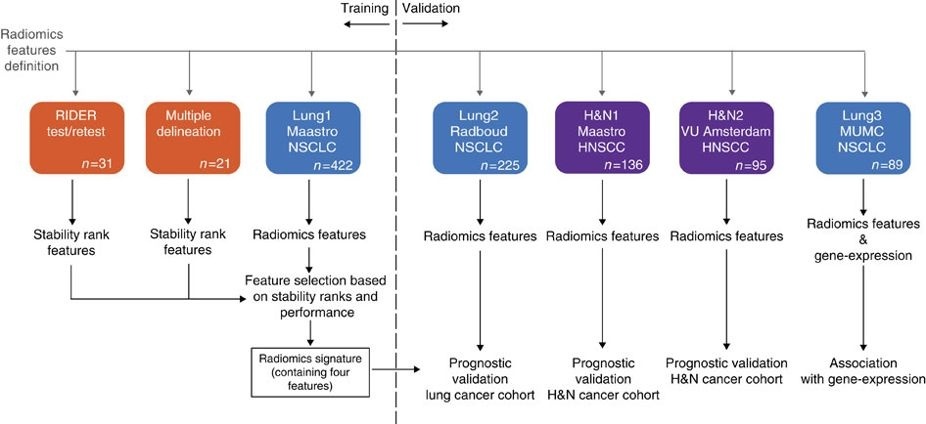
In short, this publication applies a radiomic approach to computed tomography data of 1,019 patients with lung or head-and-neck cancer. Radiomics refers to the comprehensive quantification of tumour phenotypes by applying a large number of quantitative image features. In present analysis 440 features quantifying tumour image intensity, shape and texture, were extracted. We found that a large number of radiomic features have prognostic power in independent data sets, many of which were not identified as significant before. Radiogenomics analysis revealed that a prognostic radiomic signature, capturing intra-tumour heterogeneity, was associated with underlying gene-expression patterns. These data suggest that radiomics identifies a general prognostic phenotype existing in both lung and head-and-neck cancer. This may have a clinical impact as imaging is routinely used in clinical practice, providing an unprecedented opportunity to improve decision-support in cancer treatment at low cost. The DICOM Radiotherapy Structure Sets (RTSTRUCT) and DICOM Segmentation (SEG) files in this data contain a manual delineation by a radiation oncologist of the 3D volume of the primary gross tumor volume ("GTV-1") and selected anatomical structures (i.e., lung, heart and esophagus). Of note, DICOM SEG objects contain a subset of annotations available in RTSTRUCT.
The dataset described here (Lung1) was used to build a prognostic radiomic signature. The Lung3 dataset used to investigate the association of radiomic imaging features with gene-expression profiles consisting of 89 NSCLC CT scans with outcome data can be found here: NSCLC-Radiomics-Genomics.
Other data sets in the Cancer Imaging Archive that were used in the same study published in Nature Communications: Head-Neck-Radiomics-HN1, NSCLC-Radiomics-Interobserver1, RIDER Lung CT Segmentation Labels from: Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach (RIDER-LungCT-Seg).
For scientific or other inquiries about this dataset, please contact Dr Leonard Wee (leonard.wee@maastro.nl) and Prof Andre Dekker (andre.dekker@maastro.nl) at MAASTRO Clinic/Maastricht University Medical Centre+ and Maastricht University, The Netherlandsplease contact the TCIA Helpdesk. Acknowledgements We would like to acknowledge the individuals and institutions that have provided data for this collection: - Leonard Wee, MAASTRO (Dept of Radiotherapy), Maastricht University Medical Centre+, Maastricht, Limburg, The Netherlands.
- Dirk de Ruysscher, MAASTRO (Dept of Radiotherapy), Maastricht University Medical Centre+, Maastricht, Limburg, The Netherlands.
- Andre Dekker, MAASTRO (Dept of Radiotherapy), Maastricht University Medical Centre+, Maastricht, Limburg, The Netherlands.
- Hugo Aerts, Computational Imaging and Bioinformatic Laboratory, Dana-Farber Cancer Institute & Harvard Medical School, Boston, Massachusetts, USA.
- Harmonization of the components of this dataset, including into standard DICOM representation, was supported in part by the NCI Imaging Data Commons consortium. NCI Imaging Data Commons consortium is supported by the contract number 19X037Q from Leidos Biomedical Research under Task Order HHSN26100071 from NCI.
|