Summary
The National Cancer Institute’s (NCI's) Cancer Imaging Program in collaboration with the International Society for Biomedical Imaging (ISBI) has launched a grand challenge involving prostate gland magnetic resonance imaging (MRI) data. The challenge will take place at the ISBI Symposium, April 7-11, 2013 in San Francisco, CA. |
Healthcare Context
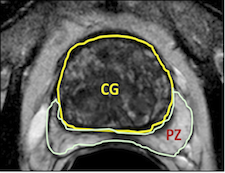
Prostate cancer is the second most common cause of cancer deaths in males and will afflict 1 in 6 men. Many diagnostic prostate biopsies fail to detect occult cancers because of the difficulty in properly determining the prostate’s anatomy. Approximately 20% of ultrasound image guided prostate biopsies return negative results despite rising levels of prostate-specific antigen (PSA) markers in the blood and the presence of cancer somewhere in the 30+ cc gland. These failures have been attributed by some to the hesitation to sample one of the two major parts of the glands— namely the predilection toward the more easily reached peripheral zone (PZ) over the less accessible central gland (CG). This year’s ISBI challenge contestants outlined these two non-overlapping adjacent regions of the gland (see image).
Advancing solutions to this vexing clinical problem will raise awareness about the impact computer image processing can have toward addressing a widespread, critical healthcare puzzle. Open and closed source algorithms applicable to both 1.5T and 3T magnetic resonance images (MRIs) will be most welcome.
Challenge Structure and Time Line
Data for the prostate challenge have been provided by Boston University and Radboud University, Nijmegen Medical Centre (the Netherlands), and divided into three groups:
- January 26: Training cases will be made publicly available to all interested participants. This consists of 60 subjects, half of which are taken from the PROSTATE-DIAGNOSIS collection (1.5T) and the other half from the Prostate-3T collection (3T). Reference Standard (or Ground Truth) data for each subject are provided in the form of NRRD image segmentations created by expert observers.
- March 7: Leader Board cases will be made available a month before the NCI-ISBI Challenge. Participants in the contest will be able to submit their segmentation results for evaluation using this data set. This will consist of 10 cases from the PROSTATE-DIAGNOSIS collection (1.5T) and the Prostate-3T collection (3T). NRRD segmentations used for generating participant scores will be held by the contest organizers.
- April 7: The 2013 NCI-ISBI Challenge will be held at the ISBI annual meeting in San Francisco, CA. Testing data will be made available to participants at this time, which will again consist of 10 cases from the PROSTATE-DIAGNOSIS collection (1.5T) and the Prostate-3T collection (3T).
Corrections/Updates:
- 3/22/13- A competitor reported an issue with case ProstateDx-01-0055, which has a dimension mismatch. The segmentation has dimensions 400x400x23 whereas the DICOM image series have dimensions of 400x400x34. We checked the case and indeed the dimensions seem to not correspond on Z (23 vs 34); however, the labels are properly spatially placed. We don't currently see a problem with using the case.
- 3/26/13- A competitor reported an issue with ProstateDx-01-0006 in which the labels for the peripheral zone and central gland were flipped. This has now been corrected in the training set NRRD zip file available below.
- 4/2/13- The testing data set is being made available in advance of the meeting for download so that participants can have it loaded prior to the meeting. The password to unlock the zip file will be provided at the ISBI meeting. See the Testing section below for full details.
Prospective participants for the Onsite Challenge Test have the option to submit a 1-page abstract to imaging.challenge@nih.gov. Please include title, authors, affiliation, objectives, data, methods, results, and conclusions. Format: 11 point font, PDF.
Obtaining the Challenge Data
Training Set
A 60-case prostate MRI 3D series "training set" consisting of Digital Imaging and Communications in Medicine (DICOM) images with matched NRRD markups that define the tissue CG and PZ are available from the NCI via The Cancer Imaging Archive (TCIA).
- Each case is ~ 3 to 10MB so you'll need to have ~600MB free on your machine.
- In order to obtain the DICOM images please login to TCIA (or register for a free account).
- Having logged on, go to the left side-bar to click on the link "search shared list".
- The Shared List Name to search for is: "ISBI Prostate Challenge - Training".
- Then click the "Submit" box at the lower right-hand part of the screen.
- Click the blue underlined link titled ISBI Prostate Challenge - Training to open the Shared List.
- For additional help with searching for shared lists please refer to section 3.7.3 in the TCIA User's Guide or contact the TCIA Helpdesk at 314-747-4254.
- Next click the blue Download Manager link.
- Be sure to note the Browse box at the top of the next screen to choose a suitable folder location on your machine to receive the downloaded DCM cases.
- Then click the start button at the bottom arrow of the screen picture below to start the download.
- Wait until all of the progress bars reach 100%. A message will appear at the bottom of the Download Manager window indicating when the data is completely finished downloading.
- For additional help with using the Download Manager please refer to section 3.8 in the TCIA User's Guide or contact the TCIA Helpdesk at 314-747-4254.
There will be 60 cases, each consisting of 15+ (4 mm thick slices at 3T) or 28+ (3 mm thick at 1.5T) adjacent axial cross-section cuts. Half of them were obtained at 1.5 T (Philips Achieva) with an endo-rectal receiver coil (fromBostonMedicalCenter) and the other half at 3T (Siemens TIM) with a surface coil (from Radboud University Nijmegen Medical Centre [RUNMC],Netherlands). They were acquired as T2-weighted MR axial pulse sequences. Each file contains nearly all DICOM acquisition parameters except tags that specifically identify Private Health Information.
To complete the training data set, each case has had individual slice CG and PZ outlines marked by Drs. Nicolas Bloch (Boston University School of Medicine) and Mirabela Rusu (Case Western University) or Drs. Henkjan Huisman, Geert Litjens, or Futterer at RUNMC Netherlands. Use the following links to download the .NRRD markups:
Leaderboard Set (released 3/7/13)
The 10 case prostate MRI 3D series "leader board set" is now available from the National Cancer Institute (NCI) via TCIA. The same instructions apply as with the training set, except for the use of a new shared list name - "ISBI Prostate Challenge - Leader Board".
- In order to obtain the DICOM images please login to TCIA (or register for a free account).
- Having logged on, go to the left side-bar to click on search shared list.
- The Shared List Name to search for is: "ISBI Prostate Challenge - Leader Board".
- Then click the Submit box at the lower right-hand part of the screen.
- Click the blue underlined link titled ISBI Prostate Challenge - Leader Board to open the Shared List.
- For additional help with searching for shared lists please refer to section 3.7.3 in the TCIA User's Guide or contact the TCIA Helpdesk at 314-747-4254.
- Next click the blue Download Manager link.
- Be sure to note the Browse box at the top of the next screen to choose a suitable folder location on your machine to receive the downloaded DCM cases.
- Then click the start button at the bottom arrow of the screen picture below to start the download.
- Wait until all of the progress bars reach 100%. A message will appear at the bottom of the Download Manager window indicating when the data is completely finished downloading.
- For additional help with using the Download Manager please refer to section 3.8 in the TCIA User's Guide or contact the TCIA Helpdesk at 314-747-4254.
Please note that no NRRD segmentations will be available for download for these leaderboard cases until after the conclusion of the challenge.
Test Set (released 4/2/13)
The 10-case prostate MRI 3D series "test set" is now available via direct download: http://challenge.kitware.com/midas/item/3046.
The file is password protected so that participants cannot use it until the day of the meeting, but will be able to download it in advance to help avoid potential issues with obtaining the data on the day of the challenge. Please note that no NRRD segmentations will be available for download for these test cases until after the conclusion of the challenge.
Submitting Your Results
Once you've trained your algorithm on those 60 cases the NCI-ISBI Challenge Committee will provide you access to a new 10-case leaderboard set to which you will not have .NRRD markup data. "Leaderboard" is a popular term (borrowed from golf) for a procedure to see how well you are doing on a pre-test set in order to get a sense of your relative competitive rank before submitting to a final test. The results of your participation in the leaderboard will be returned to you as feedback on a subset of sequestered markups retained on Kitware's MIDAS software, which analytically judges the submissions. You may then use those results for further tuning your algorithm to prepare yourself for the final test set of 10 totally new cases on which you will run your algorithm while attending the ISBI meeting in April 2013.
To submit your results to the Leader Board please visit: http://challenge.kitware.com/midas/community/5#tabs-info. Leaderboard results are also available: http://challenge.kitware.com/midas/community/5#Participant_Scoreboards.
Appendix
For a comprehensive background on the prostate, its anatomy, imaging and clinical observations see the following publications: Committee and publications that promote a PI-RADS reporting standard: For healthcare context of a sample MRI case as reported by a radiologist see linked report on training case ProstateDx-01-0001: Questions about the structure and conduct of the Challenge please contact: Questions about the TCIA image archive and downloading procedures please review the The Cancer Imaging Archive User's Guide or contact: - Phone: +1 314-747-4254
- Email: help@cancerimagingarchive.net
Questions about 3D Slicer and .nrrd file structure contact: Questions about how Kitware designed its scoring software called MIDAS see below attached document link: |
ContactsKeyvan Farahani, PhD; National Cancer Institute, NIH; (farahani@nih.gov)
Justin Kirby, BS; SAIC-Frederick; (kirbyju@mail.nih.gov)
Anant Madabhushi, PhD; Case Western Reserve University; (anant.madabhushi@case.edu)
Henkjan Huisman, PhD; Radboud University, The Netherlands; (H.Huisman@rad.umcn.nl)
Andinet Enquobahrie, PhD; Kitware Inc (andinet.enqu@kitware.com) OrganizersKeyvan Farahani, NCI
Carl Jaffe, Boston University
Nicholas Bloch, Boston University
Anant Madabhushi, Case Western Reserve University
Henkjan Huisman, Radboud University Nijmegen Medical Centre
John Freymann, SAIC-Frederick
Justin Kirby, SAIC-Frederick
Andinet Enquobahrie, Kitware Inc.
Larry Clarke, NCI |
|
|