Summary
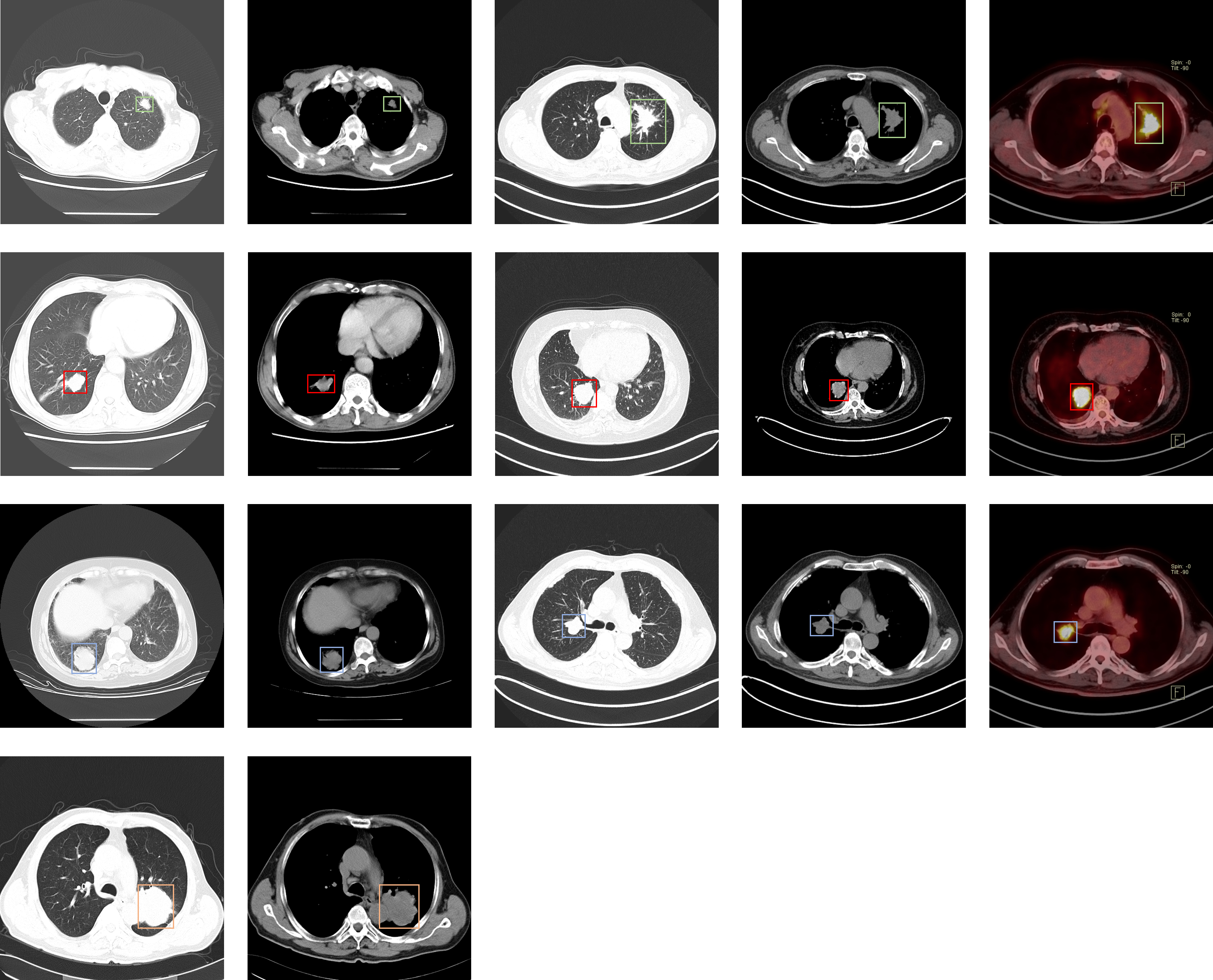
Our dataset consists of: DICOM files and XML Annotation files. The XML Annotation files, which are widely used in deep learning and machine learning research, were provided by five Radiologist (original said doctors) and two deep learning researchers making our dataset a useful tool and resource for developing algorithms for medical diagnosis. The images were analyzed on the mediastinum (window width, 350 HU; level, 40 HU) and lung (window width, 1,400 HU; level, –700 HU) settings. The reconstructions were made in 2mm-slice-thick and lung settings. The CT slice interval varies from 0.625 mm to 5 mm. Scanning mode includes plain and contrast and 3D reconstruction. The location of the tumors was labeled in the DICOM images, and the image annotations are saved in XML files in PASCAL VOC format. Users can parse the annotations using the PASCAL Development Toolkit: https://pypi.org/project/pascal-voc-tools/ Several lines about the protocol and how subjects were recruited. The cases were confirmed by pathological diagnosis. |
Acknowledgements
We would like to acknowledge the individuals and institutions that have provided data for this collection:
Drs. Huiping Han, Funing Yang and Rui Wang for their help collecting the clinical data (We aren't using the clinical data. Should this be included?)
The Computer Center and Cancer Institute at the Second Affiliated Hospital of Harbin Medical University at Harbin, Heilongjiang Province, China for their help collecting the image data
Beijing Municipal Administration of Hospital Clinical Medicine Development of Special Funding (ZYLX201511)
Hospital/Institution Name city, state, country - Special thanks to First Last Names, degree PhD, MD, etc from the Department of xxxxxx, Additional Names from same location.
- Continue with any names from additional submitting sites if collection consists of more that one.
|