Summary
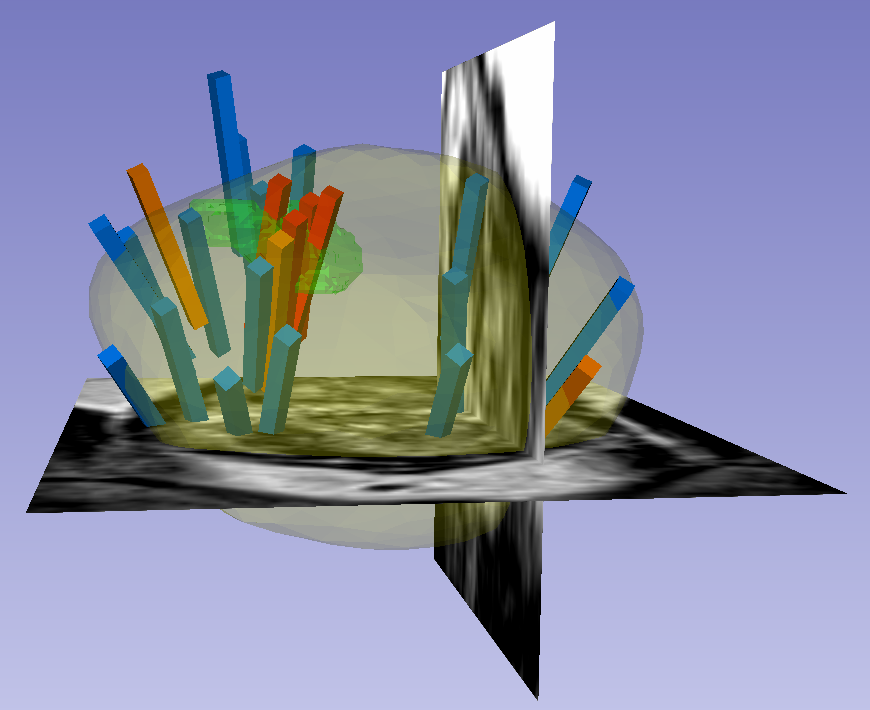
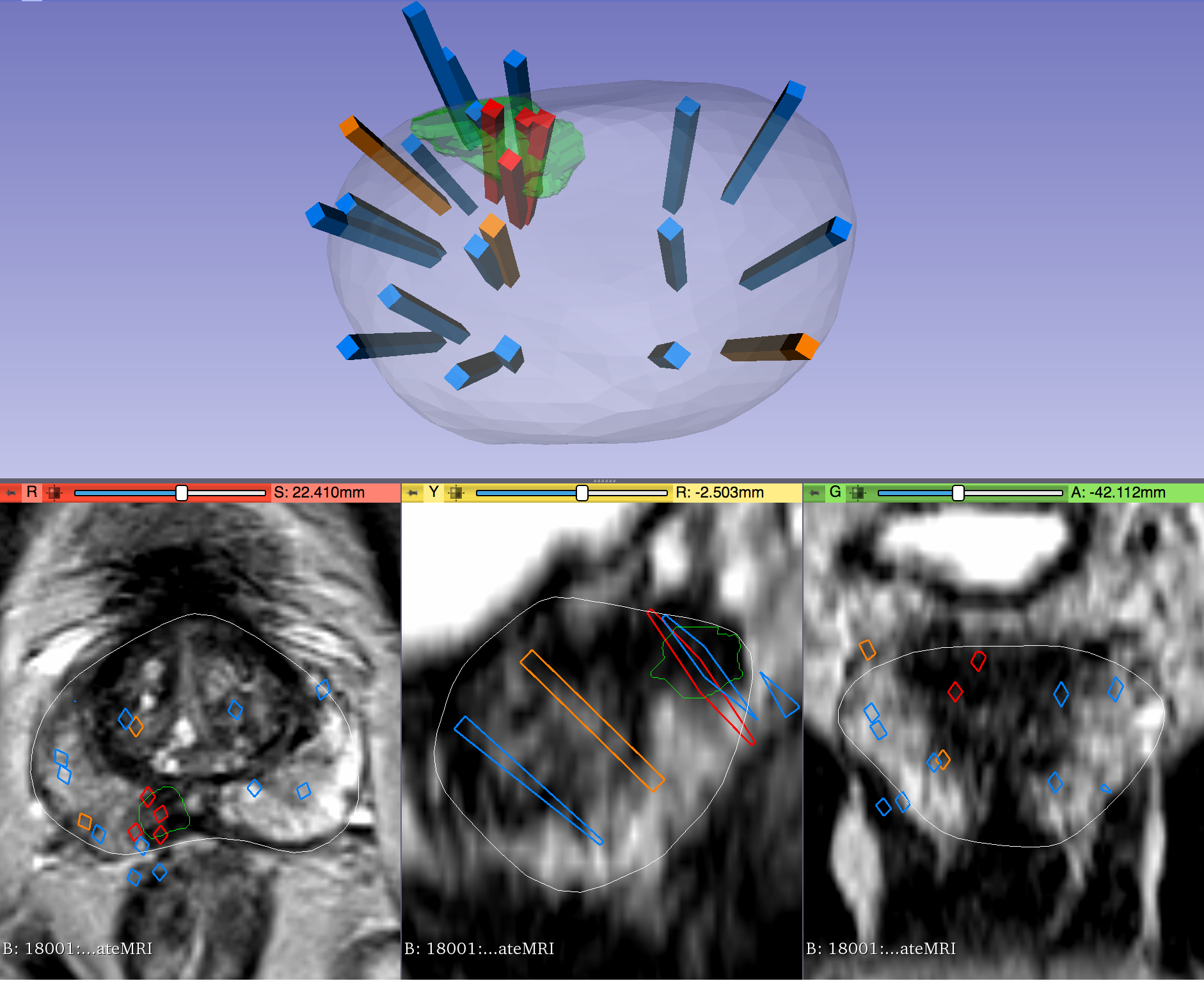
This dataset was derived from tracked biopsy sessions using the Artemis biopsy system, many of which included image fusion with MRI targets. Patients received a 3D transrectal ultrasound scan, after which nonrigid registration (e.g. “fusion”) was performed between real-time ultrasound and preoperative MRI, enabling biopsy cores to be sampled from MR regions of interest. Most cases also included sampling of systematic biopsy cores using a 12-core digital template. The Artemis system tracked targeted and systematic core locations using encoder kinematics of a mechanical arm, and recorded locations relative to the Ultrasound scan. MRI biopsy coordinates were also recorded for most cases.
MRI targets were defined using multiparametric MRI, e.g. t2-weighted, diffusion-weighted, and perfusion-weighted sequences, and scored on a Likert-like scale with close correspondence to PIRADS version 2. t2-weighted MRI was used to trace ROI contours, and is the only sequence provided in this dataset. MR imaging was performed on a 3 Tesla Trio, Verio or Skyra scanner (Siemens, Erlangen, Germany). A transabdominal phased array was used in all cases, and an endorectal coil was used in a subset of cases. The majority of pulse sequences are 3D T2:SPC, with TR/TE 2200/203, Matrix/FOV 256 × 205/14 × 14 cm, and 1.5mm slice spacing. Some cases were instead 3D T2:TSE with TR/TE 3800–5040/101, and a small minority were imported from other institutions (various T2 protocols.)
Ultrasound scans were performed with Hitachi Hi-Vision 5500 7.5 MHz or the Noblus C41V 2-10 MHz end-fire probe. 3D scans were acquired by rotation of the end-fire probe 200 degrees about its axis, and interpolating to resample the volume with isotropic resolution.
Patients with suspicion of prostate cancer due to elevated PSA and/or suspicious imaging findings were consecutively accrued. Any consented patient who underwent or had planned to receive a routine, standard-of-care prostate biopsy at the UCLA Clark Urology Center was included.
Note: Some Private Tags in this collection are critical to properly displaying the STL surface and the Prostate anatomy. Private Tag (1129,"Eigen Artemis",1016) DS VoxelSize is especially important for multi-frame US cases.
Acknowledgements
We would like to acknowledge the individuals and institutions that have provided data for this collection:
- This work was supported in part by the National Cancer Institute Award R01CA158627, Prostate Cancer SPORE at University of California-Los Angeles P50CA092131, the Beckman Coulter Foundation, Jean Perkins Foundation, and Steven C. Gordon Family Foundation
- Harmonization of the components of this dataset, including into standard DICOM representation, was supported in part by the NCI Imaging Data Commons consortium. NCI Imaging Data Commons consortium is supported by the contract number 19X037Q from Leidos Biomedical Research under Task Order HHSN26100071 from NCI
Data AccessClick the Download button to save a ".tcia" manifest file to your computer, which you must open with the NBIA Data Retriever . Click the Search button to open our Data Portal, where you can browse the data collection and/or download a subset of its contents.
| Data Type | Download all or Query/Filter |
|---|
| Images (DICOM) 77.6 (GB) | | Target Data (XLSX) 131 (KB) | | | Biopsy Data (XLSX) 4.25 (MB) | | |  (Click the "Detailed Description" tab above for usage instructions) | | Bipsy Overlays (ZIP) 333 (MB) | |
|
Detailed Description
Image Statistics |
|
|---|
Modalities | MR, US | Number of Participants | 1151 | Number of Studies | 2799 | Number of Series | 2799 | Number of Images | 61,119 | | Images Size (GB) | 77.6 |
Instructions for Proper Use of the STL Files The 3D surfaces of this dataset are in stereolithography format (binary representation). They are segmentations representing anatomic regions within the corresponding ultrasound (US) or MRI images. Each file is named "Prostate-MRI-US-Biopsy-XXXX-SURFACETYPE-seriesUID-YYYY.STL" where "XXXX" is the anonymized patient number and "YYYY" is the series instance UID of corresponding DICOM images. "SURFACETYPE" can be one of two identifiers, summarized below. - If "SURFACETYPE" is "ProstateSurface," then the STL file is a segmentation of the prostate gland. These were contoured semi-automatically with Profuse (for MRI) or the Artemis fusion biopsy system (for US), both products of Eigen (Grass Valley, CA). Using these tools, the user manually traced the prostate on one or more image planes, and the remainder of gland was automatically segmented in 3D. The user was then able to adjust vertices of the 3D segmentation as necessary. These segmentations are generally of good quality, though their accuracy is user-dependent.
- If "SURFACETYPE" is "Target," then the STL file is a segmentation of a Region of Interest (ROI) suspicious for harboring prostate cancer. Zero, one, or more than one ROI may be present, and thus are designated "Target1," "Target2," etc. The ROIs corresponding to MRI were traced manually by a radiologist based on their interpretation of the patient's multiparametric MRI study. The ROIs corresponding to US images were acquired by registering the MRI prostate surface nonrigidly and semi-automatically to the US prostate surface. Therefore, the US ROIs are subject to misregistration, with average accuracy of 3-4 mm.
- Note that proximity to an ROI increases the likelihood of, but does not guarantee, prostate cancer. Also note that radiologists assigned each ROI a Likert-like suspicion score of 1-5, with 1 corresponding to very low suspicion of prostate cancer and 5 corresponding to very high suspicion of prostate cancer. These scores, which are roughly equivalent to the "PIRADS Version 2" scoring system, can be found in a spreadsheet included with the dataset.
The STL files can be visualized in 3D Slicer as follows: 1) Import the surface(s) either by drag-and-dropping or with File->Add Data 2) Within the "Models" module, adjust Display->Opacity of the prostate surface in order to visualize any ROI(s) within 3) To visualize surfaces in the context of corresponding MRI or US images, load the corresponding image series and then toggle the Slice Display->Visible radio button within the "models" module 4) To visualize surfaces in the context of biopsy cores, import the corresponding 3D slicer file named "BiopsyVectors.mrml" by drag-and-dropping or with File->Add Data. The STL files, which are written in a standard format, can also be imported and visualized with most computer-aided design (CAD) and 3D printing software. Note that, if quantitative analysis is desired, the following transform was applied to US surfaces for 3D slicer compatibility and may need to be reversed for visualization alongside MRI or US images using other software. Transform: [0 1 0 0 0 1 1 0 0] Click the Versions tab for more info about data releases. |
Citations & Data Usage Policy
Natarajan, S., Priester, A., Margolis, D., Huang, J., & Marks, L. (2020). Prostate MRI and Ultrasound With Pathology and Coordinates of Tracked Biopsy (Prostate-MRI-US-Biopsy) [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/TCIA.2020.A61IOC1A |
Sonn, Geoffrey A., Shyam Natarajan, Daniel JA Margolis, Malu MacAiran, Patricia Lieu, Jiaoti Huang, Frederick J. Dorey, and Leonard S. Marks. "Targeted biopsy in the detection of prostate cancer using an office based magnetic resonance ultrasound fusion device." The Journal of Urology 189, no. 1 (2013): 86-92 |
Clark K, Vendt B, Smith K, Freymann J, Kirby J, Koppel P, Moore S, Phillips S, Maffitt D, Pringle M, Tarbox L, Prior F. The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository, Journal of Digital Imaging, Volume 26, Number 6, December, 2013, pp 1045-1057. DOI: 10.1007/s10278-013-9622-7 |
Other Publications Using This DataTCIA maintains a list of publications which leverage TCIA data. If you have a manuscript you'd like to add please contact the TCIA Helpdesk. |
Version 1 (Current): 2020/09/11
| Data Type | Download all or Query/Filter |
|---|
| Images (DICOM) 77.6 (GB) | | | Target Data (XLSX) 131 (KB) | | | Biopsy Data (XLSX) 4.25 (MB) | | |  (Click the "Detailed Description" tab above for usage instructions) | | Biopsy Overlays (ZIP) 333 (MB) | |
Added new subjects. |
|



