| Localtab |
|---|
| active | true |
|---|
| title | Data Access |
|---|
| Data AccessClick the Download button to save a ".tcia" manifest file to your computer, which you must open with the NBIA Data Retriever. Click the Search button to open our Data Portal, where you can browse the data collection and/or download a subset of its contents.
| Data Type | Download all or Query/Filter |
|---|
| Images (DICOM, XX.X GB) |  Image Removed Image Removed Image Removed Image Removed
| Click the Versions tab for more info about data releases. Please contact help@cancerimagingarchive.net with any questions regarding usage. | | Localtab |
|---|
| title | Detailed Description |
|---|
| | License |
|---|
| Segmentation (NIfTI, zip, 4 MB) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/DRO%20Toolkit-3%20Subjects%20SEG%20and%20QIN%20multi-site%2010%20Subjects%20SEG%20NIfTI.zip?api=v2 |
|---|
|
|
| | | Feature Variability Software Package details (xlsx, 13 kb) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Table%201%20-%20Feature%20Variability%20Software%20Details_v2.xlsx?api=v2 |
|---|
|
|
| | | DRO Results (xlsx, 31 kb) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/DRO%20Results%20Table%202%20QIN%20PET%20CT%20WG%20DRO%20Feature%20Values_Table2_supporting_data.xlsx?api=v2 |
|---|
|
|
| | | Patient Dataset Results (xlsx, 400 kb) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Patient%20Dataset%20Results%20QIN%20PET%20CT%20WG%20Patient%20Dataset%20Feature%20Values_Table3_supporting_data.xlsx?api=v2 |
|---|
|
|
| | | Harmonized GLCM Entropy Results (xlsx, 17 kb) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Harmonized%20GLCM%20Entropy%20Results%20QIN%20PET%20CT%20WG%20Patient%20Dataset%20Feature%20Values_Table4_supporting_data.xlsx?api=v2 |
|---|
|
|
| |
Collections Used in this Third Party Analysis Below is a list of the Collections used in these analyses: | Source Data Type | Download | License |
|---|
Corresponding Original CT images from LIDC-IDRI and DRO-Toolkit (DICOM, 2.0 GB) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Radiomic-Feature-Standards-DICOM%20CTs%203%20Subjects%20DRO%20Toolkit%2010%20Subjects%20QIN.tcia?api=v2 |
|---|
| Download |
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?ImageModalityCriteria=CT&MinNumberOfStudiesCriteria=1&PatientCriteria= |
|---|
|
|
Detailed Description | Image Statistics | Modalities | Number of Patients | Number of Studies | Number of Series | Number of Images | Images Size (GB) | Patient IDs for the 3 DROs from (insert DOI)
| Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-0.0-0.0 |
|
|
| ,Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-50.0-0.0 |
|
|
| ,Phantom-100.0-1.0-1.0-1.0-9.0-0.2-100.0-10.0-0.0-0.0 |
|
|
Patient IDs for the 10 LIDC-IDRI subjects (http://doi.org/10.7937/K9/TCIA.2015.LO9QL9SX)
LIDC-IDRI LIDC-IDRI-0314
LIDC-IDRI LIDC-IDRI-0325
LIDC-IDRI LIDC-IDRI-0580
LIDC-IDRI LIDC-IDRI-0766
LIDC-IDRI LIDC-IDRI-0771
LIDC-IDRI LIDC-IDRI-0811
LIDC-IDRI LIDC-IDRI-0905
LIDC-IDRI LIDC-IDRI-0963
LIDC-IDRI LIDC-IDRI-0965
LIDC-IDRI LIDC-IDRI-1012| ,LIDC-IDRI-0314,LIDC-IDRI-0325,LIDC-IDRI-0580,LIDC-IDRI-0766,LIDC-IDRI-0771,LIDC-IDRI-0811,LIDC-IDRI-0905,LIDC-IDRI-0963,LIDC-IDRI-0965,LIDC-IDRI-1012 |
| Search |
(Requires NBIA Data Retriever.) | | | Corresponding second-generation SEG images from QIN-LungCT-Seg (DICOM, 123 MB) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Radiomic-Feature-Standards-DICOM%20Segs%203%20Subjects%20DRO%20Toolkit%2010%20Subjects%20QIN.tcia?api=v2 |
|---|
| Download |
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?ImageModalityCriteria=SEG&MinNumberOfStudiesCriteria=1&PatientCriteria=Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-0.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-50.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.2-100.0-10.0-0.0-0.0,LIDC-IDRI-0314,LIDC-IDRI-0325,LIDC-IDRI-0580,LIDC-IDRI-0766,LIDC-IDRI-0771,LIDC-IDRI-0811,LIDC-IDRI-0905,LIDC-IDRI-0963,LIDC-IDRI-0965,LIDC-IDRI-1012 |
|---|
| Search |
(Requires NBIA Data Retriever.)
| |
|---|
| Localtab |
|---|
| title | Detailed Description |
|---|
| Detailed Description
DICOM Image Statistics |
|
|---|
Modalities | CT, SEG | Number of Patients | 13 | Number of Studies | 13 | Number of Series | 26 | Number of Images | 3,867 | | Images Size (GB) | 2 GB |
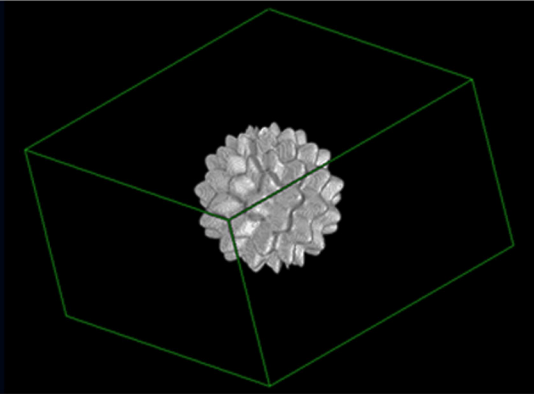
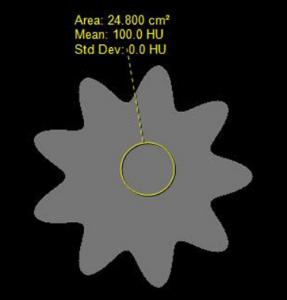
There are three datasets provided – two image datasets and one dataset consisting of three four excel spreadsheets containing feature values. - The first image dataset is a set of three Digital Reference Objects (DROs) used in the project, which are: (a) a sphere with uniform intensity, (b) a sphere with intensity variation (c) a nonspherical (but mathematically defined) object with uniform intensity. These DROs were created by the team at Stanford University and are described in (Jaggi A, Mattonen SA, McNitt-Gray M, Napel S. Stanford DRO Toolkit: digital reference objects for standardization of radiomic
AQ: F - features. Tomography. 2019;6:–.) and are a subset of the DROs described in
(link to DRO wiki page)- Stanford DRO Toolkit: Digital Reference Objects for Standardization of Radiomic Features. Each DRO is represented in both DICOM and NIfTI format and the VOI was provided in each format as well (DICOM Segmentation Object (DSO) as well as NIfTI segmentation boundary).
- The second image dataset is
a - the set of 10 patient CT scans
which is a subset of - , originating from the LIDC-IDRI dataset, that were used in the QIN multi-site collection of Lung CT data with Nodule Segmentations project (
http- https://doi.org/10.7937/K9/TCIA.2015.1BUVFJR7 )
created previously. Specifically, the same 10 cases selected from the LIDC-IDRI dataset that were used in that previous study were used in this study. As in that previous - . In that QIN study, a single lesion from each case was identified for analysis
. That previous study generated VOIs using algorithms from three academic institutions and each method performed three repeat runs on each nodule. For this study, and to - and then nine VOIs were generated using three repeat runs of three segmentation algorithms (one from each of three academic institutions) on each lesion. To eliminate one source of variability in our project, only one of the VOIs previously created for each lesion was identified and all sites used that same VOI definition. The specific VOI chosen for each lesion was the first run of the first algorithm (algorithm 1, run 1).
As in that prior project, both DICOM and NIfTI formats were created - DICOM images were provided for each
image - dataset and the VOI was provided in
each format as well (- both DICOM Segmentation Object (DSO)
as well as - and NIfTI segmentation
boundary)- formats.
- The third dataset is a collection of
three - four excel spreadsheets, each of which contains detailed information corresponding to each of the four tables in the publication. For example, the raw feature values and the summary tables for Tables 2,3 and 4 reported in the publication
below. - cited (https://doi.org/10.18383/j.tom.2019.00031). These tables are:
Software Package details : This table contains detailed information about the software packages used in the study (and listed in Table 1 in the publication) including version number and any parameters specified in the calculation of the features reported. DRO results : This contains the original feature values obtained for each software package for each DRO as well as the table summarizing results results across software packages (Table 2 in the publication) . Patient Dataset results: This contains the original feature values for each software package for each patient dataset (1 lesion per case) as well as the table summarizing results across software packages and patient datasets (Table 3 in the publication). Harmonized GLCM Entropy Results : This contains the values for the “Harmonized” GLCM Entropy feature for each patient dataset and each software package as well as the summary across software packages (Table 4 in the publication). Patient IDs for the 3 DROs from (https://doi.org/10.7937/t062-8262)
Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-0.0-0.0
Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-50.0-0.0
Phantom-100.0-1.0-1.0-1.0-9.0-0.2-100.0-10.0-0.0-0.0
Patient IDs for the 10 LIDC-IDRI subjects (https://doi.org/10.7937/K9/TCIA.2015.LO9QL9SX)
LIDC-IDRI-0314
LIDC-IDRI-0325
LIDC-IDRI-0580
LIDC-IDRI-0766
LIDC-IDRI-0771
LIDC-IDRI-0811
LIDC-IDRI-0905
LIDC-IDRI-0963
LIDC-IDRI-0965
LIDC-IDRI-1012 Additional options for download:
| DRO Data (3 subjects) | Download all or Query/Filter |
|---|
Image Data (DICOM, 452.0 MB) CT only | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/DRO%20Toolkit-3%20Subjects-DICOM-CT%20Image%20Data%20TCIA%20Manifest. |
|---|
|
|
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?ImageModalityCriteria=CT&MinNumberOfStudiesCriteria=1&PatientCriteria=Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-0.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-50.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.2-100.0-10.0-0.0-0.0&CollectionCriteria=DRO-Toolkit |
|---|
| Search |
(Requires NBIA Data Retriever.) | | Segmentation Data - DSO (DICOM, 29.0 MB) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/DRO%20Toolkit-3%20Subjects-DICOM-Segmentation%20Data%20TCIA%20Manifest.tcia?api=v2 |
|---|
| Download |
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?ImageModalityCriteria=SEG&MinNumberOfStudiesCriteria=1&PatientCriteria=Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-0.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-50.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.2-100.0-10.0-0.0-0.0&CollectionCriteria=DRO-Toolkit |
|---|
| Search |
(Requires NBIA Data Retriever.) | | Segmentation Data - (NIfTI, zip, 926 KB) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/DRO%20Toolkit-3%20Subjects-SEG%20Images%20NIfTI%20.zip?api=v2 |
|---|
| Download |
|
| Patient Datasets (10 subjects) | Download all or Query/Filter |
|---|
Image Data (DICOM, 1.0 GB) CT only |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/LIDC-IDRI-10%20Subjects-DICOM%20CT%20Image%20Data%20TCIA%20manifest.tcia?api=v2 |
|---|
| Download |
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?ImageModalityCriteria=CT&MinNumberOfStudiesCriteria=1&PatientCriteria=LIDC-IDRI-0314,LIDC-IDRI-0325,LIDC-IDRI-0580,LIDC-IDRI-0766,LIDC-IDRI-0771,LIDC-IDRI-0811,LIDC-IDRI-0905,LIDC-IDRI-0963,LIDC-IDRI-0965,LIDC-IDRI-1012&CollectionCriteria=LIDC-IDRI |
|---|
| Search |
(Requires NBIA Data Retriever.) | | Segmentation Data - (DICOM, 94 MB) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/LIDC-IDRI-10%20Subjects-DICOM%20SEG%20Image%20Data%20TCIA%20Manifest.tcia?api=v2 |
|---|
| Download |
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?saved-cart=nbia-39001586554584246 |
|---|
| Search |
(Requires NBIA Data Retriever.) | | Segmentation Data - (NIfTI, zip, 21.0 KB) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/LIDC-IDRI-10%20Subjects-NIfTI-Patient_Image_Data_NIFTI_Segs.zip?api=v2 |
|---|
| Download |
|
|
| Localtab |
|---|
| title | Citations & Data Usage Policy |
|---|
| Citations & Data Usage Policy| Tcia limited license policy |
|---|
| | Localtab |
|---|
| title | Citations & Data Usage Policy |
|---|
| Citations & Data Usage PolicyAdd any special restrictions in here. These collections are freely available to browse, download, and use for commercial, scientific and educational purposes as outlined in the Creative Commons Attribution 3.0 Unported License. Questions may be directed to help@cancerimagingarchive.net. Please be sure to acknowledge both this data set and TCIA in publications by including the following citations in your work:| Info |
|---|
| McNitt-Gray, M.*, Napel, S.*, Jaggi, A., Mattonen, S.A., Hadjiiski, L., Muzi, M., Goldgof, D., Balagurunathan, Y., Pierce, L.A., Kinahan, P.E., Jones, E.F., Nguyen, A., Virkud, A., Chan, H-P., Emaminejad, N., Wahi-Anwar, M., Daly, M., Abdalah, M., Yang, H., Lu, L., Lv, W., Rahmim, A., Gastounioti, A., Pati, S., Bakas, S., Kontos, D., Zhao, B., Kalpathy-Cramer, J., Farahani, K. (2020). Data from the Standardization in Quantitative Imaging: A Multi-center Comparison of Radiomic Features from Different Software Packages on Digital Reference Objects and Patient Datasets. Tomography, Feb, 2020.*Authors contributed equallyFeature Values [Data set]. The Cancer Imaging Archive. DOI: https://doi.org/10.7937/tcia.2020.9era-gg29. |
| Info |
|---|
| title | Publication Citation |
|---|
| Kalpathy-Cramer, J., Zhao, BMcNitt-Gray, M., Napel, S., Jaggi, A., Mattonen, S.A., Hadjiiski, L., Muzi, M., Goldgof, D., Gu, Y., Wang, XBalagurunathan, Y., Pierce, L.A., Kinahan, P.E., Jones, E.F., Nguyen, A., Virkud, A., Chan, H-P., Emaminejad, N., Wahi-Anwar, M., Daly, M., Abdalah, M., Yang, H., … Napel, S. (2016). A Comparison of Lung Nodule Segmentation Algorithms: Methods and Results from a Multi-institutional Study. Journal of Digital Imaging. Springer Nature. http., Lu, L., Lv, W., Rahmim, A., Gastounioti, A., Pati, S., Bakas, S., Kontos, D., Zhao, B., Kalpathy-Cramer, J., Farahani, K. (2020). Standardization in Quantitative Imaging: A Multi-center Comparison of Radiomic Feature Values, Tomography. https://doi.org/10.1007/s10278-016-9859-z18383/j.tom.2019.00031. |
| Info |
|---|
| Clark, K., Vendt, B., Smith, K., Freymann, J., Kirby, J., Koppel, P., Moore, S., Phillips, S., Maffitt, D., Pringle, M., Tarbox, L., & Prior, F. (2013). The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository, . Journal of Digital Imaging, Volume 26, Number (6, December, 2013, pp 1045-1057. DOI: ), 1045–1057. https://doi.org/10.1007/s10278-013-9622-7 |
| Info |
|---|
| title | Acknowledgement - Grant Citationssupport |
|---|
| - David Geffen School of Medicine at UCLA - U01CA181156
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - Stanford University School of Medicine – U01CA187947 and U24CA180927
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - University of Michigan - U01CA232931
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - University of Washington – R50CA211270, U01CA148131
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - University of South Florida - U24CA180927, U01CA200464
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - Moffitt Cancer Center – U01CA143062, U01CA200464, P30CA076292
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - UC San Francisco - U01CA225427
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - BC Cancer Research Centre - NSERC Discovery Grant: RGPIN-2019-06467
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - Columbia University- U01CA225431
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - Center for Biomedical Image Computing and Analytics at the University of Pennsylvania - U24CA189523, R01NS042645
|
| Info |
|---|
| title | Acknowledgement - Grant support |
|---|
| - Massachusetts General Hospital- U01CA154601, U24CA180927
|
In addition to the dataset citation above, please be sure to cite the following if you utilize these data in your research: | Info |
|---|
| Jayashree Kalpathy-Cramer, J., Sandy Napel, S., Dmitry Goldgof, Binsheng D., Zhao, B. (2015). Multi-site collection of Lung CT data with Nodule Segmentations. The Cancer Imaging Archive. http https://doi.org/10.7937/K9/TCIA.2015.1BUVFJR7 |
Other Publications Using This DataTCIA maintains a list of publications which leverage TCIA data. If you have a manuscript you'd like to add please contact the TCIA's Helpdesk. |
| Localtab |
|---|
| Version 1 (Current): Updated 2020/0306/XX09| Data Type | Download all or Query/Filter |
|---|
Images xxx GB) Image Removed Image Removed Image Removed Image Removed
(Requires 0 GB) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Radiomic-Feature-Standards-DICOM%20CTs%203%20Subjects%20DRO%20Toolkit%2010%20Subjects%20QIN.tcia?api=v2 |
|---|
| Download |
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?ImageModalityCriteria=CT&MinNumberOfStudiesCriteria=1&PatientCriteria=Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-0.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-50.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.2-100.0-10.0-0.0-0.0,LIDC-IDRI-0314,LIDC-IDRI-0325,LIDC-IDRI-0580,LIDC-IDRI-0766,LIDC-IDRI-0771,LIDC-IDRI-0811,LIDC-IDRI-0905,LIDC-IDRI-0963,LIDC-IDRI-0965,LIDC-IDRI-1012 |
|---|
| Search |
(Requires NBIA Data Retriever.) | | Corresponding second-generation SEG images from QIN-LungCT-Seg (DICOM, 123 MB) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Radiomic-Feature-Standards-DICOM%20Segs%203%20Subjects%20DRO%20Toolkit%2010%20Subjects%20QIN.tcia?api=v2 |
|---|
| Download |
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?ImageModalityCriteria=SEG&MinNumberOfStudiesCriteria=1&PatientCriteria=Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-0.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.0-100.0-10.0-50.0-0.0,Phantom-100.0-1.0-1.0-1.0-9.0-0.2-100.0-10.0-0.0-0.0,LIDC-IDRI-0314,LIDC-IDRI-0325,LIDC-IDRI-0580,LIDC-IDRI-0766,LIDC-IDRI-0771,LIDC-IDRI-0811,LIDC-IDRI-0905,LIDC-IDRI-0963,LIDC-IDRI-0965,LIDC-IDRI-1012 |
|---|
| Search |
(Requires |
Segmentations (NIfTI, XX.X GB) | | | Segmentation (NIfTI, zip, 4 MB) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/DRO%20Toolkit-3%20Subjects%20SEG%20and%20QIN%20multi-site%2010%20Subjects%20SEG%20NIfTI.zip?api=v2 |
|---|
|
|
| | Feature Variability Software Package details (xlsx) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Table%201%20-%20Feature%20Variability%20Software%20Details_v2.xlsx?api=v2 |
|---|
|
|
| | DRO Results (xlsx) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/DRO%20Results%20Table%202%20QIN%20PET%20CT%20WG%20DRO%20Feature%20Values_Table2_supporting_data.xlsx?api=v2 |
|---|
|
|
| | Patient Dataset Results (xlsx) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Patient%20Dataset%20Results%20QIN%20PET%20CT%20WG%20Patient%20Dataset%20Feature%20Values_Table3_supporting_data.xlsx?api=v2 |
|---|
|
|
| | Harmonized GLCM Entropy Results (xlsx) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/70222123/Harmonized%20GLCM%20Entropy%20Results%20QIN%20PET%20CT%20WG%20Patient%20Dataset%20Feature%20Values_Table4_supporting_data.xlsx?api=v2 |
|---|
|
|
 Image Removed Image Removed
|
|