...
In addition we would like to list these publications here on our web site. If you have utilized TCIA in your research please contact us at CancerImagingArchive@mail.nih.gov so that we can include your publications in the list below. The publication list below includes references to the original data collection as well as publications that specifically used data from TCIA. You can obtain the publications specifically based on TCIA in Endnote xml format. This should be usable as input to your favorite reference management system.
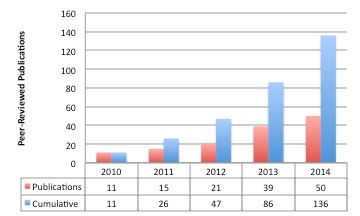
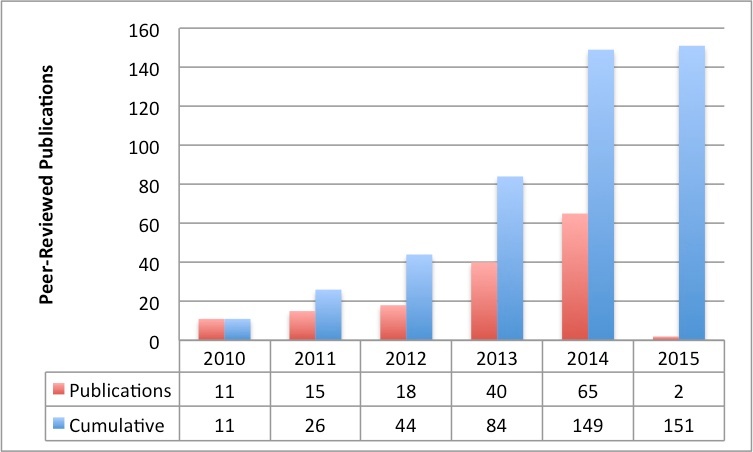
TCIA-Related Publication History
...
- Fedorov A, Fluckiger J, Ayers GD, Li X, Gupta SN, Tempany C, Mulkern R, Yankeelov TE, Fennessy FM. A comparison of two methods for estimating DCE-MRI parameters via individual and cohort based AIFs in prostate cancer: A step towards practical implementation. Magnetic resonance imaging. 2014;32(4):321-9.
- Hegde JV, Mulkern RV, Panych LP, Fennessy FM, Fedorov A, Maier SE, Tempany C. Multiparametric MRI of prostate cancer: An update on state‐of‐the‐art techniques and their performance in detecting and localizing prostate cancer. Journal of Magnetic Resonance Imaging. 2013;37(5):1035-54.
Collection: QIN Sarcoma
- Meyer JM, Perlewitz KS, Hayden JB, Doung Y-C, Hung AY, Vetto JT, Pommier RF, Mansoor A, Beckett BR, Tudorica A. Phase I trial of preoperative chemoradiation plus sorafenib for high-risk extremity soft tissue sarcomas with dynamic contrast-enhanced MRI correlates. Clinical Cancer Research. 2013;19(24):6902-11.
...
Collection: Radiogenomics
Pope WB. Genomics of Brain Tumor Imaging. Neuroimaging Clinics of North America. 2015;25(1):105-19.
- Colen R, Foster I, Gatenby R, Giger ME, Gillies R, Gutman D, Heller M, Jain R, Madabhushi A, Madhavan S, Napel S, Rao A, Saltz J, Tatum J, Verhaak R, Whitman G. NCI Workshop Report: Clinical and Computational Requirements for Correlating Imaging Phenotypes with Genomics Signatures. Translational Oncology. 2014;7(5):556-69. doi: http://dx.doi.org/10.1016/j.tranon.2014.07.007.
- Rao A. Exploring relationships between multivariate radiological phenotypes and genetic features: A case-study in Glioblastoma using the Cancer Genome Atlas, Global Conference on Signal and Information Processing (GlobalSIP), 2013 IEEE.
- Hunter L. Radiomics of NSCLC: Quantitative CT Image Feature Characterization and Tumor Shrinkage Prediction. Thesis, University of Texas; 2013.
- Karnayana PM. Radiogenomic correlation for prognosis in patients with glioblastoma multiformae. Journal of Neuro-Oncology. 2013 June 13 (link to thesis)
...
- Natteshan N, Jothi JAA. Automatic Classification of Brain MRI Images Using SVM and Neural Network Classifiers. Advances in Intelligent Informatics. Springer. 2015 320:19-30. doi:10.1007/978-3-319-11218-3_3 (link)Colen RR, Vangel M,
Wangaryattawanich P, Wang J, Thomas GA, Chaddad A, Zinn PO, Colen RR, editors. Survival analysis of pre-operative GBM patients by using quantitative image features. Control, Decision and Information Technologies (CoDIT), 2014 International Conference on; 2014: IEEE.
Colen RR, Wang J, Singh SK, Gutman DA, Zinn PO. Glioblastoma: Imaging Genomic Mapping Reveals Sex-specific Oncogenic Associations of Cell Death. Radiology. 2014.
- Colen RR, Vangel M, Wang J, Gutman DA, Hwang SN, Wintermark M, Rajan J, Jilwan-Nicola M, Gutman DA, Hwang SN, Wintermark M, Rajan J, Jilwan-Nicola M, Chen JY, Raghavan P, Holder CA, Rubin D, Huang E, Kirby J, Freymann J, Jaffee CC, Flanders A, Zinn PO. Imaging genomic mapping of an invasive MRI phenotype predicts patient outcome and metabolic dysfunction: a TCGA glioma phenotype research group project.BMC Medical Genomics, 2014. 7(1):30. doi:10.1186/1755-8794-7-30 (link)
- Gevaert O, Mitchell LA, Achrol AS, Xu J, Echegaray S, Steinberg GK, Chesier SH, Napel S, Zaharchuk G, Plevritis SK. Glioblastoma Multiforme: Exploratory Radiogenomic Analysis by Using Quantitative Image Features. Radiology, 2014. doi: 10.1148/radiol.14131731 (link)
- Mazurowski MA, Zhang J, Peters KB, and Hobbs H. Computer-extracted MR imaging features are associated with survival in glioblastoma patients. Journal of Neuro-Oncology. 2014 Aug 24 [Epub ahead of print] doi: 10.1007/s11060-014-1580-5 (link)
- Jain R, Poisson L, Gutman D, Scarpace L, Hwang SN, Holder C, Wintermark M, Colen RR, Kirby J, Freymann J, Jaffe C, Mikkelsen T, Flanders A. Outcome Prediction in Patients with Glioblastoma by Using Imaging, Clinical, and Genomic Biomarkers: Focus on the Nonenhancing Component of the Tumor. Radiology. 2014 Aug;272(2):484-93. doi: 10.1148/radiol.14131691. Epub 2014 Mar 19. 2014 (link)
- Nicolasjilwan M, Hu Y, Yan C, Meerzaman D, Holder CA, Gutman D, et al. Addition of MR imaging features and genetic biomarkers strengthens glioblastoma survival prediction in TCGA patients. Journal of Neuroradiology, July 2014. doi: 10.1016/j.neurad.2014.02.006
- Wassal E, Zinn P, Colen R. DIFFUSION AND CONVENTIONAL MR IMAGING GENOMIC BIOMARKER SIGNATURE FOR EGFR MUTATION IDENTIFICATION IN GLIOBLASTOMA. Neuro-Oncology. 2014;16(suppl 5):v156-v7.
- Wassal E, Zinn P, Colen R. DIFFUSION AND CONVENTIONAL MR IMAGING GENOMIC BIOMARKER SIGNATURE PREDICTS IDH-1 MUTATION IN GLIOBLASTOMA PATIENTS. Neuro-Oncology. 2014;16(suppl 5):v157-v.
Steed T, Treiber J, Patel K, Taich Z, White N, Treiber M, Farid N, Carter B, Dale A, Chen C. Iterative Probabilistic Voxel Labeling: Automated Segmentation for Analysis of The Cancer Imaging Archive Glioblastoma Images. American Journal of Neuroradiology. 2014.
Kwon D, Shinohara RT, Akbari H, Davatzikos C. Combining Generative Models for Multifocal Glioma Segmentation and Registration. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2014: Springer; 2014. p. 763-70.
- Amer A, Zinn P, Colen R. IMMEDIATE POST OPERATIVE VOLUME OF ABNORMAL FLAIR SIGNAL PREDICTS PATIENT SURVIVAL IN GLIOBLASTOMA PATIENTS. Neuro-Oncology. 2014;16(suppl 5):v138-v.
- Amer A, Zinn P, Colen R. IMMEDIATE POST-RESECTION PERICAVITARIAN DWI HYPERINTENSITY IN GLIOBLASTOMA PATIENTS IS PREDICTIVE OF PATIENT OUTCOME. Neuro-Oncology. 2014;16(suppl 5):v138-v9.
- Gutman DA, Cooper LAD, Hwang SN, Holder CA, Gao J, Aurora TD, Dunn WD, Scarpace L, Mikkelsen T, Jain R, Wintermark M, Jilwan M, Raghavan P, Huang E, Clifford RJ, Monqkolwat P, Kleper V, Freymann J, Kirby J, Zinn PO, Moreno CS, Jaffe C, Colen R, Rubin DL, Saltz J, Flanders A, Brat DJ. MR Imaging Predictors of Molecular Profile and Survival: Multi-institutional Study of the TCGA Glioblastoma Data Set. Radiology. 2013 May:267(2):560-569,doi:10.1148/radiol.13120118 (link)
- Jain R, Poisson L, Narang J, Gutman D, Scarpace L, Hwang SN, Holder C, Wintermark M, Colen RR, Kirby J, Freymann J, Brat DJ, Jaffe C, Mikkelsen T. Genomic Mapping and Survival Prediction in Glioblastoma: Molecular Subclassification Strengthened by Hemodynamic Imaging Biomarkers. Radiology, 2013 Apr:267(1):212 –220, doi:10.1148/radiol.12120846 (link)
- Mazurowski MA, Desjardins A, Malof JM. Imaging descriptors improve the predictive power of survival models for glioblastoma patients. Neuro-oncology, 2013. 15(10):1389-1394 (link)
- Zinn PO, Colen RR. Imaging Genomic Mapping in Glioblastoma. Neurosurgery 60:126-130. Aug 2013 (link)
- Jain R, Poisson L, Narang J, Scarpace L, Rosenblum ML, Rempel S, Mikkelson T. Correlation of Perfusion Parameters with Genes Related to Angiogenesis Regulation in Glioblastoma: A Feasibility Study. American Journal of Neuroradiology, 2012. 33(7):1343-1348 [Epub ahead of print] (link)
- Zinn PO, Sathyan P, Mahajan B, Bruyere J, Hegi M, et al. (2012) A Novel Volume-Age-KPS (VAK) Glioblastoma Classification Identifies a Prognostic Cognate microRNA-Gene Signature. PLoS ONE, 2012 7(8): e41522. doi:10.1371/journal.pone.0041522 (link)
- Zinn PO, Majadan B, Sathyan P, Singh SK, Majumder S, et al. 2011Radiogenomic Mapping of Edema/Cellular Invasion MRI-Phenotypes in Glioblastoma Multiforme. PLoS ONE, 2011 6(10): e25451. doi:10.1371/journal.pone.0025451 (link)
...
Collection: Algorithm Development
- Blessy SPS, Sulochana CH. Performance analysis of unsupervised optimal fuzzy clustering algorithm for MRI brain tumor segmentation. Technology and Health Care. 2014.
- ElNawasany AM, Ali AF, Waheed ME. A Novel Hybrid Perceptron Neural Network Algorithm for Classifying Breast MRI Tumors. Advanced Machine Learning Technologies and Applications: Springer; 2014. p. 357-66.
- Hong S, Huang Y, Cao Y, Chen X, Han J-DJ. Approaches to uncovering cancer diagnostic and prognostic molecular signatures. Molecular & Cellular Oncology. 2014.
- Codella N, Connell J, Pankanti S, Merler M, and Smith JR. Automated Medical Image Modality Recognition by Fusion of Visual and Text Information. Medical Image Computing and Computer-Assisted Intervention. 2014, Springer. 487-495. (link)
- Ertugrul OF. Adaptive Texture Energy Measure Method. International Journal of Intelligent Information Systems. 2014. 3(2):13-18. doi:10.11648/j.ijiis.20140302.11 (link)
- Kawa J, Juszczyk J, Pyciński B, Badura P, Pietka E. Radiological Atlas for Patient Specific Model Generation. Information Technologies in Biomedicine, 2014 4:69-82. 10.1007/978-3-319-06596-0_7. (link)
- Kowalik-Urbaniak I, Brunet D, Wang J, Koff D, Smolarski-Koff N, Vrscay ER, Wallace B, Wang Z.The quest for ‘diagnostically lossless’ medical image compression: a comparative study of objective quality metrics for compressed medical images. SPIE Medical Imaging. 2014. Vol. 9073. International Society for Optics and Photonics. doi:10.1117/12.2043196 (link)
- Naresh P and Shettar R. Image Processing and Classification Techniques for Early Detection of Lung Cancer for Preventive Health Care: A Survey. International Journal of Recent Trends in Engineering & Technology, 2014. 11:595-601 (link)
- Patel NP, Parmar SK, and Jain KR. Swift Pre Rendering Volumetric Visualization of Magnetic Resonance Cardiac Images based on Isosurface Technique. Procedia Technology, 2014. 14:422-429. DOI: 10.1016/j.protcy.2014.08.054 (link)
- Roy S, Brown MS, and Shih GL. Visual Interpretation with Three-Dimensional Annotations (VITA): Three-Dimensional Image Interpretation Tool for Radiological Reporting. Journal of Digital Imaging, 2014. 27(1):49-57. doi: 10.1007/s10278-013-9624-5 (link)
- Sivakumar S, and Chandrasekar C. A Study on Image Denoising for Lung CT Scan Images.International Journal of Emerging Technologies in Computational and Applied Sciences, 2014. 7(1):86-91 (link)
- Harmon S, Wendelberger B, and Jeraj R. SU-E-J-98: Radiogenomics: Correspondence Between Imaging and Genetic Features Based On Clustering Analysis. Medical Physics, 2014. 41(6): p. 178-178. doi:http://dx.doi.org/10.1118/1.4888150 (link)
- Krishnakumar V. and Parthiban L. Performance Analysis of Denoising in MR Images with Double Density Dual Tree Complex Wavelets, Curvelets and NonSubsampled Contourlet Transforms. Annual Review & Research in Biology, 2014. 4(19):2938-2956. doi:10.9734/ARRB/2014/9131#sthash.qFePVdL1.dpuf (link)
- Codella N, Merler M. IBM TJ Watson Research Center. Semantic Model Vector for ImageCLEF2013. June 18, 2014. (link)
- Agostinelli F, Anderson MR, and Lee H. Adaptive Multi-Column Deep Neural Networks with Application to Robust Image Denoising. Advances in Neural Information Processing Systems. 2013. (link)
Agostinelli F, Anderson MR, Lee H, editors. Robust Image Denoising with Multi-Column Deep Neural Networks. Advances in Neural Information Processing Systems; 2013.
- Breseman K, Lee C, Bloch BN, and Jaffe C. Constructing 3D-Printable CAD Models of Prostates from MR Images. Bioengineering Conference (NEBEC),
39th Annual Northeast , IEEE, 27-28. 5-7 April 2013. doi:10.1109/NEBEC.2013.8 - Buckler A, Liu TT, Savig E, Suzek BE, Rubin DL, and Paik D. Quantitative Imaging Biomarker Ontology (QIBO) for Knowledge Representation of Biomedical Imaging Biomarkers. Journal of Digital Imaging, 2013. 26(4):630-641. doi:10.1007/s10278-013-9599-2 (link)
- Heyns M, Breseman K, Lee C, Bloch BN, Jaffe C, and Xiang H. Design of a Patient-Specific Radiotherapy Treatment Target. Bioengineering Conference (NEBEC), 2013 39th Annual Northeast. 2013.171-172. IEEE.doi:10.1109/NEBEC.2013.75
- Kumar A, Kim J, Cai W, Fulham M, and Feng D. Content-Based Medical Image Retrieval: A Survey of Applications to Multidimensional and Multimodality Data. Journal of Digital Imaging, 2013. 26(6):1025-1039. doi: 10.1007/s10278-013-9619-2.(link)
- Lundström C. vPSNR: a visualization-aware image fidelity metric tailored for diagnostic imaging. International Journal of Computer Assisted Radiology and Surgery, 2013. 8(3):437-450. doi: 10.1007/s11548-012-0792-4 (link)
- Olmedo I, Guerra Perez Y, Johnson JF, Raut L, Hoe DHK. Image segmentation on GPGPUs: a cellular automata-based approach. Proceedings of the 2013 Summer Computer Simulation Conference. Society for Modeling & Simulation International. 2013. 51. (link)
- Pambrun JF, Noumeir R. Compressibility variations of JPEG2000 compressed computed tomography. Conference Proceedings, 35th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, 2013:3375-3378. doi: 10.1109/EMBC.2013.6610265 (link)
- Roozgard A, Barzigar N, Verma P, and Cheng S. 3D medical image denoising using 3D block matching and low-rank matrix completion. Signals, Systems and Computers, Asilomar Conference, 3-6 Nov. 2013, 253 – 257 IEEE. doi:10.1109/ACSSC.2013.6810271
- Yankeelov TE, Atuegwu N, Hormuth D, et al. Clinically Relevant Modeling of Tumor Growth and Treatment Response. Sci Transl Med. 2013 May 29;5(187):187ps9 doi: 10.1126/scitranslmed.3005686 (link)
- Huang L-C, Tseng L-Y, Hwang M-S. A reversible data hiding method by histogram shifting in high quality medical images. Journal of Systems and Software. 2013;86(3):716-27. doi: http://dx.doi.org/10.1016/j.jss.2012.11.024.
- Huang LC, Yseng LY, Hwang MS. A reversible data hiding method by histogram shifting in high quality medical images. Journal of Systems and Software 2013 March;86(3):716-27 doi: 10.1016/j.jss.2012.11.024 (link)
- Pheng HS and Shamsuddin SM. Texture classification of lung computed tomography images. 2012 International Conference on Graphic and Image Processing. 2013. Vol. 8768. International Society for Optics and Photonics. doi:10.1117/12.2011108 (link)
- Barzigar N, Roozgard A, Verma P, Cheng S. Removing Mixture Noise from Medical Images Using Block Matching Filtering and Low-Rank Matrix Completion. Healthcare Informatics, Imaging and Systems Biology, IEEE International Conference. 2012.134. doi:10.1109/HISB.2012.59 (link)
- Otake Y, Schafer S, Stayman JW, Zbijewski W, Kleinszig G, Graumann R, Khanna AJ, Siewerdsen JH. Automatic localization of target vertebrae in spine surgery using fast CT-to-fluoroscopy (3D-2D) image registration. SPIE Medical Imaging, 2012. Volume: 8316. International Society for Optics and Photonics. doi:10.1117/12.911308 (link)
- Roozgard A, Cheng AS, Liu H. Malignant nodule detection on lung ct scan images with kernel rx-algorithm. Biomedical and Health Informatics (BHI), 2012 IEEE-EMBS International Conference on 5-7 Jan. 2012 499 – 502. IEEE. doi: 10.1109/BHI.2012.6211627.
- Biancardi AM, Jirapatnakul AC, Reeves AP. A comparison of ground truth estimation methods. International Journal of Computer Assisted Radiology and Surgery, 2010. 5(3):295-305. DOI: 10.1007/s11548-009-0401-3 (link)
- Soysal OM, Chen P, Schneider H. An Image Processing Tool for Efficient Feature Extraction in Computer-Aided Detection Systems. Granular Computing (GrC) IEEE International Conference 2010. 14-16 Aug. 438-442. doi:10.1109/GrC.2010.128
- Tseng LY and Huang LC. Automatic fissure detection in CT images based on the genetic algorithm. Machine Learning and Cybernetics (ICMLC), International Conference. IEEE. 2010. 5: 2583 – 2588. DOI:10.1109/ICMLC.2010.5580871
...