| Localtab |
|---|
| active | true |
|---|
| title | Data Access |
|---|
| Data AccessClick the Download button to save a ".tcia" manifest file to your computer, which you must open with the
NBIA Retriever . Click the Search button to open our Data Portal, where you can browse the data collection and/or download a subset of its contents.Data | Type | Download all or Query/Filter | License |
|---|
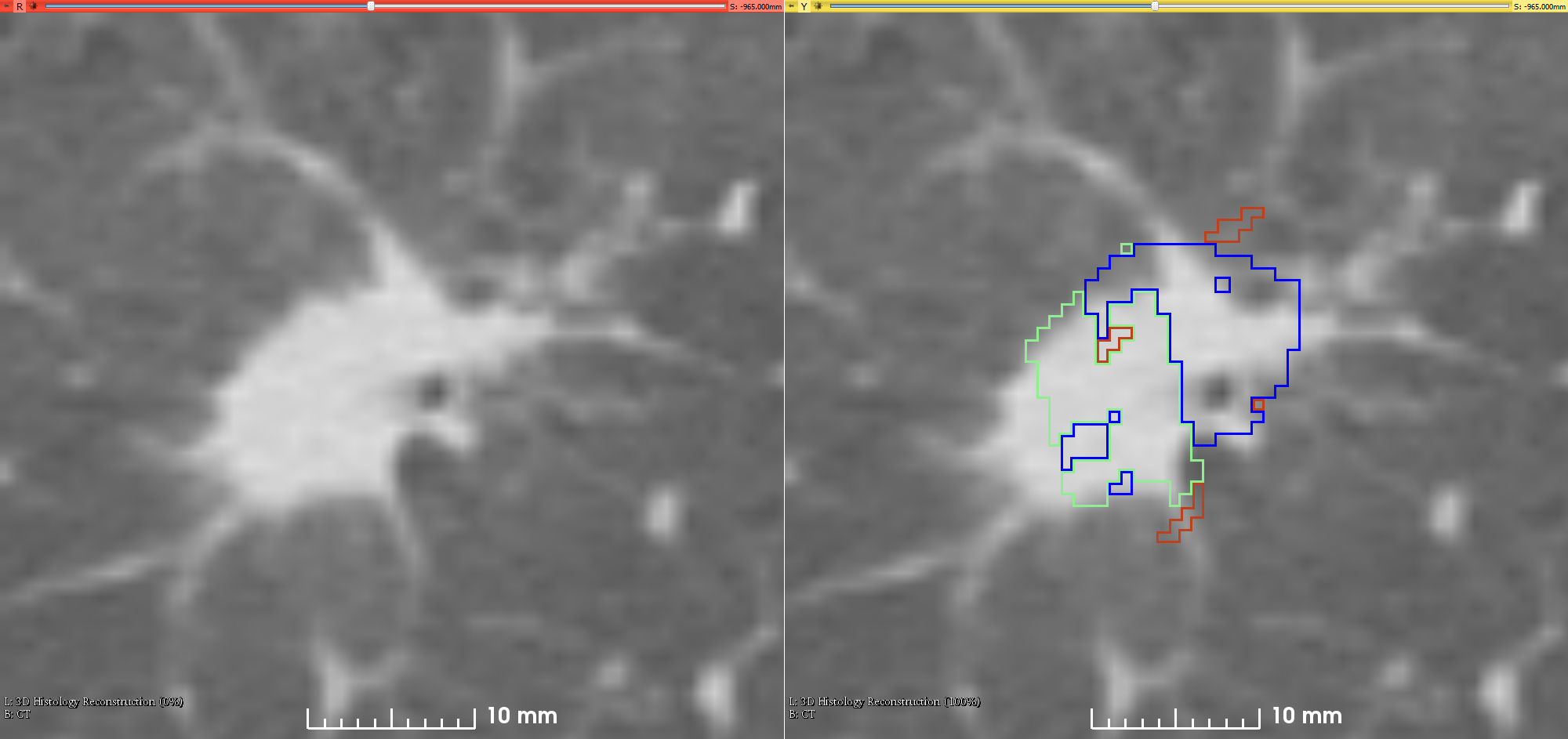
| CT Images and Histology compartments mapped on CT (DICOM, 5.5 GB) |
 Image Removed Image Removed  Image Removed Image Removed |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/39878702/TCIA_Lung-Fused-CT-Pathology-2018-07-30.tcia?version=1&modificationDate=1541510138720&api=v2 |
|---|
|
|
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?CollectionCriteria=Lung-Fused-CT-Pathology |
|---|
|
|
(Download requires the NBIA Data Retriever) | | | Annotated Whole Slide Pathology Images (TIF, 21.3 GB) |
| Tcia button generator |
|---|
| url | https://faspex.cancerimagingarchive.net/aspera/faspex?context=eyJyZXNvdXJjZSI6InBhY2thZ2VzIiwidHlwZSI6ImV4dGVybmFsX2Rvd25sb2FkX3BhY2thZ2UiLCJpZCI6IjUyNSIsInBhc3Njb2RlIjoiYjIyM2M1M2RjZjc2ODYzODFlYTRjMWM1MjliNTcwMmQyOGE3YjgwOSIsInBhY2thZ2VfaWQiOiI1MjUiLCJlbWFpbCI6ImhlbHBAY2FuY2VyaW1hZ2luZ2FyY2hpdmUubmV0In0= |
|---|
|
|
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://pathdb.cancerimagingarchive.net/imagesearch?f[0]=collection:lung_fused_ct_pathology |
|---|
|
|
(Download and apply the IBM-Aspera-Connect plugin to your browser to retrieve this faspex package) |
 Image Removed Image Removed | | | Clinical data (XLSX, 5 kB) |
 Image Removed Image Removed |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/39878702/Lung%20Fused-CT-Pathology%20spreadsheet.xlsx?version=1&modificationDate=1532544176723&api=v2 |
|---|
|
|
| |
Click the Versions tab for more info about data releases. | Nci_crdc additional resources |
|---|
The following external resources have been made available by the data submitters. These are not hosted or supported by TCIA, but may be useful to researchers utilizing this collection. |
| Localtab |
|---|
| title | Detailed Description |
|---|
| Detailed Description
| | Pathology Image Statistics |
|---|
Modalities | CT, Histology compartments mapped on CT (DICOM) | Pathology | Number of Participants | 6 | 6 | Number of Studies | 636 | 6 | Number of Series | 52 | N/A | Number of Images | 11,210 | 25 | | Image Size (GB) | 5.5 | 21.3 |
Supporting DocumentationWithin the directory CT_Segmentations_and_annotations/CT_Segmentations/<ptID>/ there are five directories: - BloodVessels (derived from CT)
- Nodule (derived from CT)
- MappedFromHistologyBloodVessels
- MappedFromHistologyInvasion
- MappedFromHistologyLesion
The data set is fully described in the following publications: Rusu et al. Co-registration of pre-operative CT with ex vivo surgically excised ground glass nodules to define spatial extent of invasive adenocarcinoma on in vivo imaging: a proof-of-concept study. European Radiology (2018); PMCID:PMC5630490 DOI:10.1007/s00330-017-4813-0 Histology Data DescriptionThere is one folder for each patient, with the same folder name as the TCIA ID. Each folder with TCIA ID name contains 2 folders: - “images”
- Contains scanned histology images (in tiff format) pertinent to the TCIA ID Patient, e.g. LungFCP-01-0001_b1.tiff, LungFCP-01-0001_b2.tiff, etc
- “annotations”
- Contains the pathologist’s annotations, stored as tiff file with the same image size as the scanned histology files, with 0 where there is not label, and 255 where a region was annotated. It is possible to have multiple annotations for each file, e.g. for file LungFCP-01-0001\images\LungFCP-01-0001_b1.tiff the following regions are available:
- LungFCP-01-0001_b1_annotation_00_R000G255B000.tiff
- LungFCP-01-0001_b1_annotation_00_R255G000B000.tiff
A part of the filenames (LungFCP-01-0001_b1_annotation_00_RxxxGyyyBzzz) indicate the type of annotation: - Adenocarcinoma in Situ : R000G000B255, R001G000B255, R002G000B255
- Invasive Adenocarcinoma: R000G255B000, R001G255B000, R002G255B000, R003G255B000, R004G255B000, R005G255B000, R006G255B000
- Invasive Adenocarcinoma + Adenocarcinoma in Situ: R255G000B000
If multiple files are available for the same type of annotation, e.g. Invasive Adenocarcinoma, it indicates that the pathologists has annotated multiple regions Description Content of the content of “FinalPublishedResults” folderThe folder “FinalPublishedResults” contains one folder for each patient. Within each patient folder, there are 3 subfolders: “CT”, ”Histology”, & ”Results_XX”. When expanded each patient folder appears as depicted below. - LungFCP-01-0001
- LungFCP-01-0002
- LungFCP-01-0003
- LungFCP-01-0004
- LungFCP-01-0005
- LungFCP-01-0006
The folder “CT” contains the 3D CT volume saved as one mha file, the segmentation of the lesion obtained by majority voting for the three radiologist segmentations, and the segmentation of the blood vessels. The folder “histology” has a similar content as the raw histology data, but a lower resolution, with some additional processing, e.g. applied gross rotation and flipping to correct for artifacts related to the mounting on the glass slide, and with additional annotations beside the lesion and in situ disease, e.g. blood vessels. The folder Results contain the outcome of the registration of the histopathology images and the CT. All scripts used for generating these results are available at https://github.com/mirabelarusu/RadPathFusionLung Please refer to the following publication for the methodological details: Rusu M., Rajiah P., Gilkeson R., Yang M., Donatelli C., Thawani R., Jacono F.J., Linden P., Madabushi A. (2017) Co-registration of pre-operative CT with ex vivo surgically excised ground glass nodules to define spatial extent of invasive adenocarcinoma on in vivo imaging: a proof-of-concept study. European Radiology 27:10, 4209:4217. DOI: https://doi.org/10.1007/s00330-017-4813-0 |
| Localtab |
|---|
| title | Citations & Data Usage Policy |
|---|
| Citations & Data Usage Policy | publictcia-collectionlimited-license-policy |
|---|
| Info |
|---|
| title | Publication Citation |
|---|
| Rusu et al. , M., Rajiah, P., Gilkeson, R., Yang, M., Donatelli, C., Thawani, R., Jacono, F. J., Linden, P., & Madabhushi, A. (2018) Co-registration of pre-operative CT with ex vivo surgically excised ground glass nodules to define spatial extent of invasive adenocarcinoma on in vivo imaging: a proof-of-concept study. European Radiology (2018); PMCIDVol. 27, Issue 10, pp. 4209–4217). PMCID:PMC5630490 DOI: https://doi.org/10.1007/s00330-017-4813-0 |
| Info |
|---|
| Clark K, Vendt B, Smith K, Freymann J, Kirby J, Koppel P, Moore S, Phillips S, Maffitt D, Pringle M, Tarbox L, Prior F. The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository, Journal of Digital Imaging, Volume 26, Number 6, December, 2013, pp 1045-1057. (paper). DOI: https://doi.org/10.1007/s10278-013-9622-7 |
Additional Publication ResourcesThe Collection authors suggest the below will give context to this dataset: - Rusu M., Rajiah P., Gilkeson R., Yang M., Donatelli C., Thawani R., Jacono F.J., Linden P., Madabushi A. (2017) Co-registration of pre-operative CT with ex vivo surgically excised ground glass nodules to define spatial extent of invasive adenocarcinoma on in vivo imaging: a proof-of-concept study. European Radiology 27:10, 4209:4217. DOI: https://doi.org/10.1007/s00330-017-4813-0
Other Publications Using This DataTCIA maintains a list of publications that which leverage our data. At this time we are not aware of any publications based on this data. If you have a publication you'd like to add, please contact the TCIA's Helpdesk. |
| Localtab |
|---|
| Version 1 (Current) Updated 2018/07-/30-2018
| Data Type | Download all or Query/Filter |
|---|
| Images (DICOM, 5.5 GB) | | Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/39878702/TCIA_Lung-Fused-CT-Pathology-2018-07-30.tcia?version=1&modificationDate=1541510138720&api=v2 |
|---|
|
|
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://nbia.cancerimagingarchive.net/nbia-search/?CollectionCriteria=Lung-Fused-CT-Pathology |
|---|
|
|
 Image Removed Image Removed  Image Removed Image Removed (Requires the NBIA Data Retriever .)
| Annotated Whole Slide Pathology Images (TIF, 21.3 GB)  Image Removed Image Removed |
| Tcia button generator |
|---|
| url | https://faspex.cancerimagingarchive.net/aspera/faspex?context=eyJyZXNvdXJjZSI6InBhY2thZ2VzIiwidHlwZSI6ImV4dGVybmFsX2Rvd25sb2FkX3BhY2thZ2UiLCJpZCI6IjUyNSIsInBhc3Njb2RlIjoiYjIyM2M1M2RjZjc2ODYzODFlYTRjMWM1MjliNTcwMmQyOGE3YjgwOSIsInBhY2thZ2VfaWQiOiI1MjUiLCJlbWFpbCI6ImhlbHBAY2FuY2VyaW1hZ2luZ2FyY2hpdmUubmV0In0= |
|---|
|
|
| Tcia button generator |
|---|
| label | Search |
|---|
| url | https://pathdb.cancerimagingarchive.net/imagesearch?f[0]=collection:lung_fused_ct_pathology |
|---|
|
|
(Download and apply the IBM-Aspera-Connect plugin to your browser to retrieve this faspex package) | | Clinical data (XLSX) |
| Tcia button generator |
|---|
| url | https://wiki.cancerimagingarchive.net/download/attachments/39878702/Lung%20Fused-CT-Pathology%20spreadsheet.xlsx?version=1&modificationDate=1532544176723&api=v2 |
|---|
|
 Image Removed Image Removed
|
|
|