| Redirect | ||||
|---|---|---|---|---|
|
Summary
| Excerpt |
|---|
The Burdenko 'sGlioblastoma Progression Dataset ( B-GBM-PDBGPD) is a systematic data collection of pre-radiotherapy MRI images offrom 180 patients with primary glioblastoma treated at the Burdenko National Medical Research Center of Neurosurgery between 2014 and 2020. We provide four MRI sequences for all patients - T1, T1c
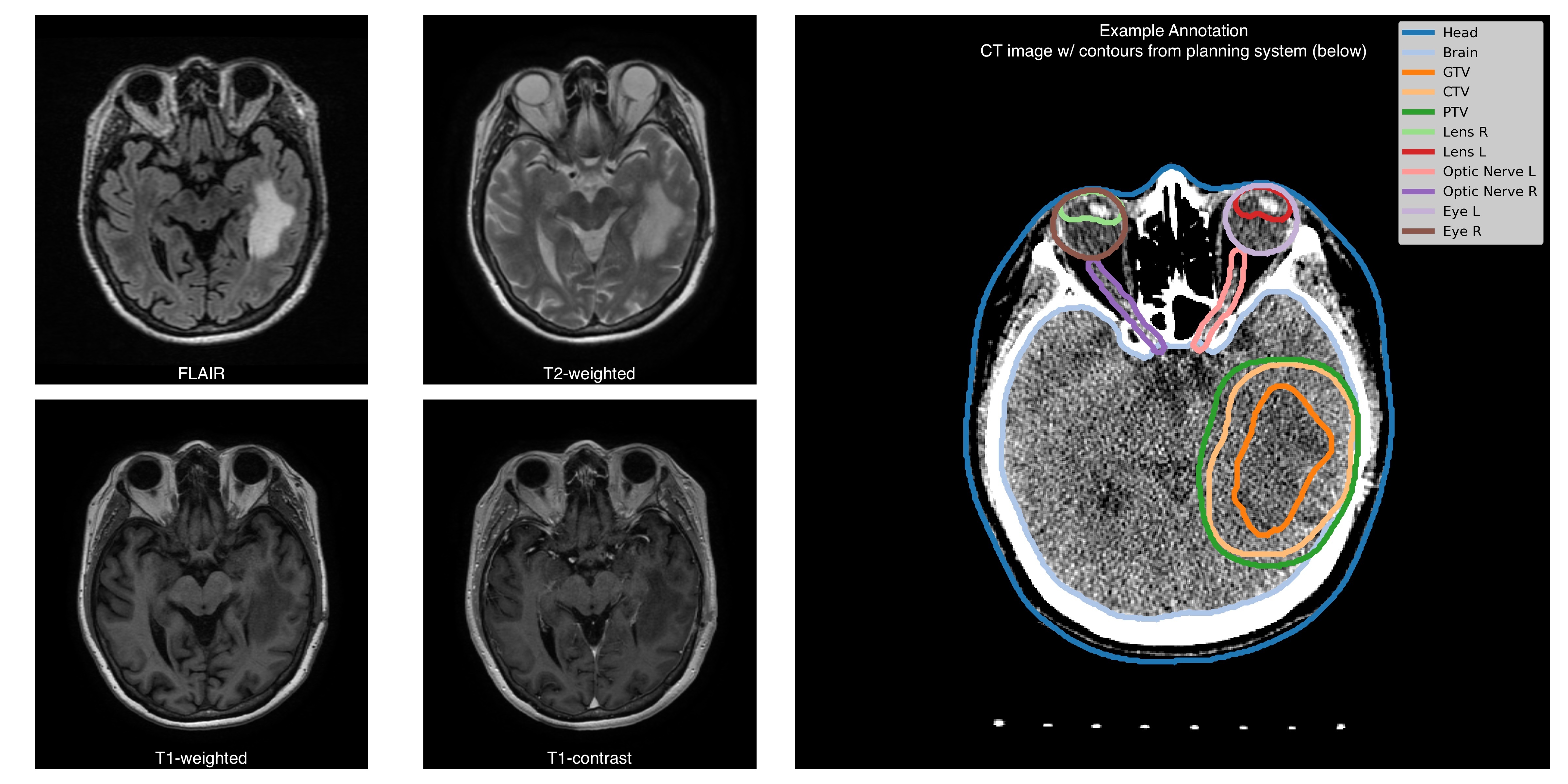
For each patient, the dataset includes imaging studies conducted for radiotherapy planning and follow-up studies. The radiotherapy studies consist of 4 MRI sequences (T1, T1C, T2, FLAIR ;), a topometric CT scan ; and annotations of the GTV area made for radiotherapy planning. Additionally, the data collection contains follow, and associated radiotherapy planning files (RTSTRUCT, RTPLAN, and RTDOSE). Follow-up studies (from 1 to 78-time pointsper patient) . Each time-point includesinclude 2-4 MRI sequences (with a minimal set of T1c,T1C and FLAIR) per patient ) with contours of the irradiated volumes. The RTDOSE, RTPLAN files; additional. Additional genetic information (IDH1/2, MGMT mutations); andand a treatment response status ( tumortumour progression, tumortumour pseudoprogression, treatment response) are available for a subset of subjectspatients. The MRI studies were performed on variousobtained from different sites, with scanners from different4 vendors and varying scanning protocols.CT studies were conducted in Burdenko National Medical Research Center of Neurosurgery on a single scanner.Dataset was collected in the radiation therapy department. The benchmark on the current dataset might be closer to clinical practice owing to data minimal preprocessing, heterogeneity in scanning modalities, sufficient size of the dataset, and longitudinal patient imaging data. It is a novel dataset compared to other TCIA collections. It contains four MRI imaging sequences for each patient (and a CT scan) and manual annotations of the glioblastoma gross tumor volumes (GTV) for radiation treatment planning, thus, complementing BraTS and TCIA-GBM datasets for brain tumor segmentation. The task of delineating GTV currently lacks large publicly available data. Availability of follow-up images allows for assessment of treatment response and development of disease progression models. We provide biological and clinical information along with imaging data: sex, age, idh1/2, mgmt mutations, and treatment responseobtained using a unified scanning protocol. Planning and follow-up studiesEach RTSTRUCT contains information on Gross Tumour Volume (GTV), Clinical Target Volume (CTV), and Planning Target Volume (PTV). In addition, each RTSTRUCT includes annotations of 10 anatomical structures: Eyes (Left, Right), Lenses (Left, Right), Optic Nerves (Left, Right), Brain, Brain stem, Chiasm, and external contour (Head). For a subset of patients RTSTRUCTs include annotation of Gross Tumour Volume assessed on follow-up (longitudinal) studies. For each patient, we supplement imaging information with clinical data: IDH1/2 gene mutation is available for 110 patients (97 negative, 13 positive), MGMT promoter methylation status is available for 92 patients (55 negative, 37 positive), death date is available for 80 patients (anonymized with preserved time shift), surgery date (anonymized with preserved time shift) and treatment response status (treatment response, tumour progression, tumour pseudoprogression) are available for all control studies. Additional informationAll MRIs are provided in the original acquisition space, while RTSTRUCTs, RTPLANs, and RTDOSEs are aligned with topometric CT scans. MRI to CT registration files are not provided as a part of the collection, however, here is a github link supporting a containerized solution (written in Python, based on ANTs library) that runs all necessary images and masks alignment. A subset of MRI images are para-transversal (direction cosine vectors as stored in Image Orientation Patient DICOM attribute form an orthonormal basis, but not a canonical one). |
Acknowledgements
We would like to acknowledge the individuals and institutions that have provided data for this collection:
...
Hospital/Institution Name city, state, country - Special thanks to First Last Names, degree PhD, MD, etc from the Department of xxxxxx, Additional Names from same location.
...
- Department of Radiosurgery and Radiotherapy of the Burdenko National Medical Research Center of Neurosurgery staff who were involved in the preparation of this dataset.
- Special thanks to Stanislav Krasnyanskiy, MSc, and Gennady Gorlachev, Ph.D. for the technical support of the data export.
| Localtab Group | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|