Versions Compared
Key
- This line was added.
- This line was removed.
- Formatting was changed.
| Info | ||
|---|---|---|
| ||
Le Hou, Rajarsi Gupta, John S. Van Arnam, Yuwei Zhang, Kaustubh Sivalenka, Dimitris Samaras, Tahsin M. Kurc, Joel H. Saltz (2019). Dataset of Segmented Nuclei in Hematoxylin and Eosin Stained Histopathology Images of 10 Cancer Types [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/tcia.2019.4a4dkp9u |
Description
Large dataset of nucleus segmentations in whole slide tissue images with quality control results are available here. There are two subsets of data: (1) automatic nucleus segmentation data of 5,060 whole slide tissue images of 10 cancer types, with quality control results. (2) manual nucleus segmentation data of 1,356 image patches from the same 10 cancer types plus additional 4 cancer types.
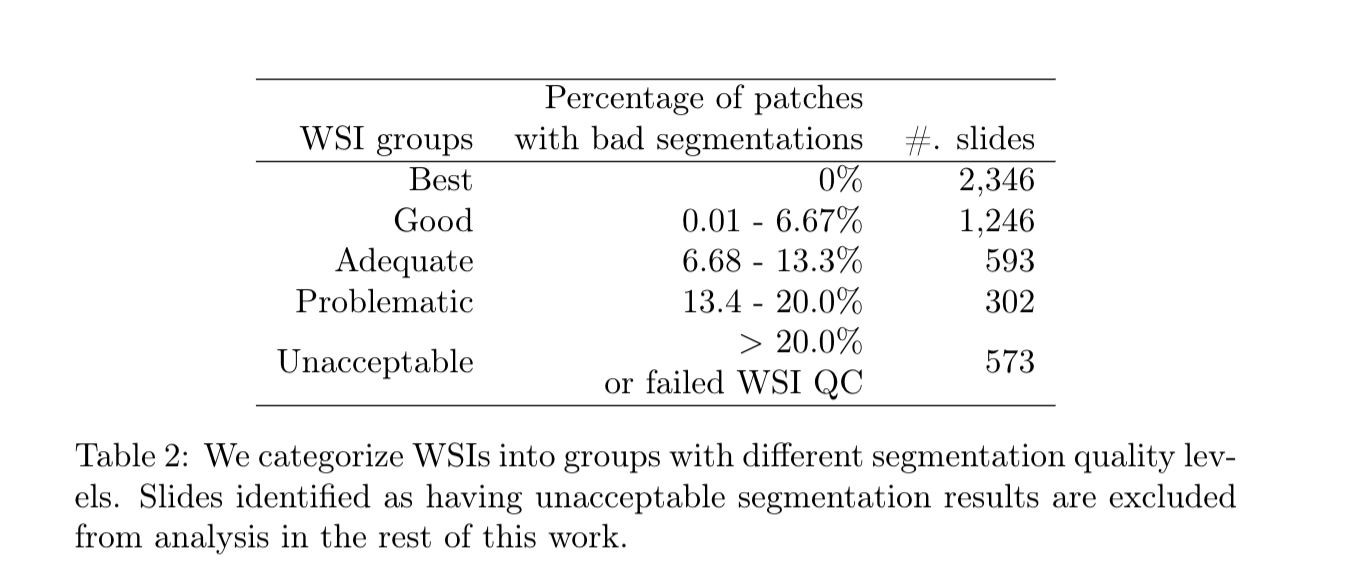
These 5,060 Whole Slide Images (WSIs) are from the following 10 cancer types:
BLCABladder urothelial carcinoma
BRCABreast invasive carcinoma
CESCCervical squamous cell carcinoma and endocervical adenocarcinoma
GBMGlioblastoma Multiforme
LUADLung adenocarcinoma
LUSCLung squamous cell carcinoma
PAADPancreatic adenocarcinoma
PRADProstate adenocarcinoma
SKCMSkin Cutaneous Melanoma
UCECUterine Corpus Endometrial Carcinoma
Note that you can also download segmentation data of following 4 cancer types, although they are not officially verified or released.
COAD Colon adenocarcinoma
READ Rectal adenocarcinoma
STAD Stomach adenocarcinoma
UVM Uveal Melanoma
| Info | ||
|---|---|---|
| ||
Hou, Le, Ayush Agarwal, Dimitris Samaras, Tahsin M. Kurc, Rajarsi R. Gupta, and Joel H. Saltz. "Robust Histopathology Image Analysis: To Label or to Synthesize?." In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 8533-8542. 2019. Open Access Here |
Download
Note: Please contact help@cancerimagingarchive.net with any questions regarding usage.
- List of histopathology slides
- Unacceptable results are marked with “qa_wsi_failed” or “qa_patch_failed”.
- WSI quality control result
- Random segmentation region checking result
- Automatic nucleus segmentation data
- Visualize segmentation data on PathDB here
Manual nucleus segmentation data of 1,356 patches
These 1,356 patches are randomly extracted from all 14 cancer types mentioned above. This data contains original H&E stained histopathology image patches, and instance-level segmentation masks. Additional information is in the readme.txt file of this data. Download here
| HTML |
|---|
<p xmlns:dct="http://purl.org/dc/terms/" xmlns:vcard="http://www.w3.org/2001/vcard-rdf/3.0#">
<a rel="license"
href="http://creativecommons.org/publicdomain/zero/1.0/">
<img src="http://i.creativecommons.org/p/zero/1.0/88x31.png" style="border-style: none;" alt="CC0" />
</a>
<br />
To the extent possible under law,
<a rel="dct:publisher"
href="https://doi.org/10.7937/tcia.2019.4a4dkp9u">https://doi.org/10.7937/tcia.2019.4a4dkp9u</a>
has waived all copyright and related or neighboring rights to
<span property="dct:title">Dataset of Segmented Nuclei in Hematoxylin and Eosin Stained Histopathology Images of 10 Cancer Types (collection nucleus:segmentation)</span>.
This work is published from:
<span property="vcard:Country" datatype="dct:ISO3166"
content="US" about="https://doi.org/10.7937/tcia.2019.4a4dkp9u">
United States</span>.
</p> |