When data is submitted to TCIA it undergoes an extensive curation process to assure completeness, proper formatting to facilitate discovery and data reuse and removal of all protected health information. Once data is released on the public TCIA repository it is Published to the world. This publication is associated with the creation of a Digital Object Identifier that allows direct access to the data.
In addition to data publication via TCIA we strongly urge researchers who submit data to TCIA to also submit a Data Descriptor publication to a journal such as Nature Scientific Data. In this type of publication the authors will describe the data acquisition process, the experiment that drove this data collection and value of the data for future research (see each journal for specific content requirements). A Data Descriptor is a scientific paper that includes the DOI to the data previously published on TCIA and helps to call the attention of the scientific community to the data you have submitted. The details provided in a Data Descriptor publication greatly enhance the value of your contribution.
A Data Descriptor is different from a scholarly paper in which you describe your experiment and present the results of your analysis. Many journals do not provide sufficient space for details of data acquisition. So today you can provide those details and the data you collected by making full use of TCIA and journals that support data publication. In summary we urge you to:
Please remember in all of your publications based on TCIA data to include appropriate references to TCIA so we can identify your publications, reference them, and make them easily available to other researchers from the TCIA web site. These citations are critical for providing continued justification of funding from the agencies that support TCIA, and are what allow us to provide this data to you free of charge. Guidelines for how to cite TCIA can be found on our Citation Guidelines wiki page. In addition we would like to list these publications here on our web site. If you have utilized TCIA in your research please contact us at help@cancerimagingarchive.net so that we can include your publications in the list below. The publication list below includes references to the original data collection as well as publications that specifically used data from TCIA.
Download citation list (Endnote XML format)
For convenience you can obtain the publications specifically based on TCIA in Endnote XML format:Pubs_basedon_TCIA_0617.xml. This should be usable as input to your favorite reference management system.
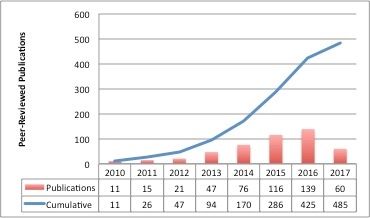
TCIA-Related Publication History

Table of Contents
Prior, F., Smith, K., Sharma, A., Kirby, J., Tarbox, L., Clark, K., Bennett, W., Nolan, T., Freymann, J. (2017). The public cancer radiology imaging collections of The Cancer Imaging Archive. Nature Scientific Data, 4; 1-7. doi:10.1038/sdata.2017.124
Kohli, M., Morrison, J. J., Wawira, J., Morgan, M. B., & Hostetter, J., Genereaux, B., Hussain, M., Langer S. G. (2017). Creation and curation of the society of imaging informatics in medicine hackathon dataset. Journal of Digital Imaging, 1-4. doi:10.1007/s10278-017-0003-5
Parks, C.L., Monson, K.L. (2016). Automated Facial Recognition of Computed Tomography-Derived Facial Images: Patient Privacy Implications. Journal of Digital Imaging. 1-11. DOI: 10.1007/s10278-016-9932-7
Huang, B.E., Mulyasasmita, W., Rajagopal, G. (2016). The Path from Big Data to Precision Medicine. Expert Review of Precision Medicine and Drug Development, 1(2):129-143. (link)
Freymann JB, Kirby JS, Perry JH, Clunie DA, and Jaffe CC. Image data sharing for biomedical research—meeting HIPAA requirements for de-identification.Journal of Digital Imaging 25, no. 1 (2012): 14-24. (paper)
Lehrer, M., Bhadra, A., Ravikumar, V., Chen, J. Y., Wintermark, M., Hwang, S. N., Holder, C. A., Huang, E. P., Fevrier-Sullivan, B., Freymann, J. B., Rao, A., & TCGA Glioma Phenotype Research Group. (2017). Multiple-response regression analysis links magnetic resonance imaging features to de-regulated protein expression and pathway activity in lower grade glioma. Oncoscience, 4, 57-66. doi:10.18632/oncoscience.353
Liu TT, Achrol AS, Mitchell LA, Rodriguez SA, Feroze A, Iv M, Kim C, Chaudhary N, Gevaert O, Stuart JM, Harsh GR, Chang SD, Rubin DL. Magnetic resonance perfusion image features uncover an angiogenic subgroup of glioblastoma patients with poor survival and better response to antiangiogenic treatment. Neuro-Oncology. 2016:1-11. doi: 10.1093/neuonc/now270
Schrock M, Batar B, Lee J, Druck T, Ferguson B, Cho J, Akakpo K, Hagrass H, Heerema N, Xia F. Wwox–Brca1 interaction: role in DNA repair pathway choice. Oncogene. 2016:1-13. doi: 10.1038/onc.2016.389.
Song SE, Bae MS, Chang JM, Cho N, Ryu HS, Moon WK. MR and mammographic imaging features of HER2-positive breast cancers according to hormone receptor status: a retrospective comparative study. Acta Radiologica. 2016:0284185116673119.
McCann SM, Jiang Y, Fan X, Wang J, et al. Quantitative Multiparametric MRI Features and PTEN Expression of Peripheral Zone Prostate Cancer: A Pilot Study. AJR Am J Roentgenol (2016). 206(3):559-565 (link)
Katrib A, Hsu W, Bui A, Xing Y. “Radiotranscriptomics”: A synergy of imaging and transcriptomics in clinical assessment. Quantitative Biology. 2016:1-12. (link)
Zhu, Y., H. Li, et al. (2015). TU-CD-BRB-06: Deciphering Genomic Underpinnings of Quantitative MRI-Based Radiomic Phenotypes of Invasive Breast Carcinoma. Medical physics 42(6): 3603-3603.
Tomczak K, Czerwińska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (Pozn). 2015;19(1A):A68-A77.
Pope WB. Genomics of Brain Tumor Imaging. Neuroimaging Clinics of North America. 2015;25(1):105-19.
Feldman, M., M. G. Piazza, et al. (2015). 137 Somatostatin Receptor Expression on VHL-Associated Hemangioblastomas Offers Novel Therapeutic Target. Neurosurgery 62: 209-210.
Gevaert O, Xu J, Hoang CD, Leung AN, Xu Y, Quon A, Rubin DL, Napel S, Plevritis SK. Non-small cell lung cancer: identifying prognostic imaging biomarkers by leveraging public gene expression microarray data--methods and preliminary results. Radiology. 2012;264(2):387-96. Epub 2012/06/23. (link)
Vani, N., Swomya, A., & Jayamma, N. (2017). Brain tumor classification using support vector machine. International Research Journal of Engineering and Technology, 1724-1729. (link)
Kaur T, Saini BS, Gupta S. A joint intensity and edge magnitude-based multilevel thresholding algorithm for the automatic segmentation of pathological MR brain images. Neural Computing and Applications. 2016:1-24. doi: 10.1007/s00521-016-2751-4
Song J, Liu Z, Zhong W, Huang Y, Ma Z, Dong D, Liang C, Tian J. Non-small cell lung cancer: quantitative phenotypic analysis of CT images as a potential marker of prognosis. Scientific reports. 2016;6:38282:1-9. doi: 10.1038/srep38282
Crawford L, Monod A, Chen AX, Mukherjee S, Rabadán R. Topological Summaries of Tumor Images Improve Prediction of Disease Free Survival in Glioblastoma Multiforme. arXiv preprint arXiv:161106818. 2016:1-29.
Zheng C, Wang X, Feng D, editors. Topology guided demons registration with local rigidity preservation. Engineering in Medicine and Biology Society (EMBC), 2016 IEEE 38th Annual International Conference; 2016: IEEE.
Kotrotsou A, Zinn PO, Colen RR. Radiomics in Brain Tumors: An Emerging Technique for Characterization of Tumor Environment. Magnetic Resonance Imaging Clinics of North America. 2016;24(4):719-29.
Parmar, C., R. T. Leijenaar, et al. (2015). "Radiomic feature clusters and Prognostic Signatures specific for Lung and Head &Neck cancer." Sci Rep 5: 11044.
Dhara AK, Mukhopadhyay S, Alam N, Khandelwal N. Measurement of spiculation index in 3D for solitary pulmonary nodules in volumetric lung CT images. Proc. SPIE 8670, Medical Imaging 2013: Computer-Aided Diagnosis, 86700K. (link)
Hsieh KL-C, Tsai R-J, Teng Y-C, Lo C-M. Effect of a computer-aided diagnosis system on radiologists' performance in grading gliomas with MRI. PloS one. 2017;12(2):e0171342 (link)
Hsieh KL-C, Lo C-M, Hsiao C-J. Computer-aided grading of gliomas based on local and global MRI features. Computer Methods and Programs in Biomedicine. 2017;139:31-8. DOI: doi.org/10.1016/j.cmpb.2016.10.021
Yang H, Liu F, Wang Z, Tang H, Sun S, Sun S. Research on the Content-Based Classification of Medical Image. Journal of Medical Imaging and Health Informatics. 2017;7(1):129-36. (link)
Rezaie AA, Habiboghli A. Detection of Lung Nodules on Medical Images by the Use of Fractal Segmentation. International Journal of Interactive Multimedia and Artificial Inteligence. 2017;4(Special Issue on 3D Medicine and Artificial Intelligence):15-9. (link)
Chen H, Zhang Y, Zhang W, Liao P, Li K, Zhou J, Wang G. Low-dose CT via convolutional neural network. Biomedical Optics Express. 2017;8(2):679-94.(link)
Roth HR, Lu L, Seff A, Cherry KM, Hoffman J, Wang S, Liu J, Turkbey E, Summers RM. A new 2.5 D representation for lymph node detection using random sets of deep convolutional neural network observations. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2014: Springer; 2014. p. 520-7.
Seff A, Lu L, Cherry KM, Roth HR, Liu J, Wang S, Hoffman J, Turkbey EB, Summers RM. 2d view aggregation for lymph node detection using a shallow hierarchy of linear classifiers. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2014: Springer; 2014. p. 544-52.
Agostinelli F, Anderson MR, Lee H, editors. Robust Image Denoising with Multi-Column Deep Neural Networks. Advances in Neural Information Processing Systems; 2013.
Kumar, D., A. Wong, et al. (2015). Lung Nodule Classification Using Deep Features in CT Images. Computer and Robot Vision (CRV), 2015 12th Conference on, IEEE.
Kanas, V. G., E. I. Zacharaki, et al. (2015). "A low cost approach for brain tumor segmentation based on intensity modeling and 3D Random Walker." Biomedical Signal Processing and Control 22: 19-30.
Magdy, E., N. Zayed, et al. (2015). "Automatic Classification of Normal and Cancer Lung CT Images Using Multiscale AM-FM Features." International Journal of Biomedical Imaging 2015.
Zayed, N. and H. A. Elnemr (2015). "Statistical Analysis of Haralick Texture Features to Discriminate Lung Abnormalities." International Journal of Biomedical Imaging 2015.
Jaffray D, Chung C, Coolens C, Foltz W, Keller H, Menard C, Milosevic M, Publicover J, Yeung I, editors. Quantitative imaging in radiation oncology: An emerging science and clinical service. Seminars in Radiation Oncology; 2015: Elsevier.
Androutsou, T. Clinical Decision Support System for Lung Cancer Diagnosis by analysis of thoracic CT images. Carrier NTUA, Department of Electrical and Computer Engineering 2017. (link to thesis)
Albalooshi FA. Self-organizing Approach to Learn a Level-set Function for Object Segmentation in Complex Background Environments. University of Dayton; 2015. (link to thesis)
Camlica Z. Image Area Reduction for Efficient Medical Image Retrieval. Waterloo, Ontario, Canada,: University of Waterloo; 2015. (link to thesis)
Karnayana PM. Radiogenomic correlation for prognosis in patients with glioblastoma multiformae. San Diego State University; 2013. (link to thesis)
Nabizadeh, N. Automated Brain Lesion Detection and Segmentation Using Magnetic Resonance Images. Electrical and Computer Engineering. Miami, FL, University of Miami. PhD., 2015. (link to thesis)
Wieser, H.-P. Supervised Machine Learning Approach Utilizing Artificial Neural Networks for Automated Prostate Zone Segmentation in Abdominal MR images. Klagenfurt, Austria, Fachhochschule Kärnten/Carinthia University of Applied Sciences; 2013.(link to thesis)
Gevaert O, Mitchell LA, et al. (2014). Glioblastoma multiforme: exploratory radiogenomic analysis by using quantitative image features. TCIA. Saint Louis, MO. (link)
Gutman DA, Cooper LA, et al. (2014). MR Imaging Predictors of Molecular Profile and Survival: Multi-institutional Study of the TCGA Glioblastoma Data Set. TCIA. Saint Louis, MO. (link)
Huang W, Li X, et al. (2014). Variations of dynamic contrast-enhanced magnetic resonance imaging in evaluation of breast cancer therapy response: a multicenter data analysis challenge. TCIA. Saint Louis, MO. (link)
Jain R, Poisson LM, et al. (2014). Outcome Prediction in Patients with Glioblastoma by Using Imaging, Clinical, and Genomic Biomarkers: Focus on the Nonenhancing Component of the Tumor. TCIA. Saint Louis, MO. (link)
Kalpathy-Cramer J, Napel S, et al. (2015). QIN multi-site collection of Lung CT data with Nodule Segmentations. TCIA. Saint Louis, MO. (link)
Messay T, Hardie RC, et al. (2014). Segmentation of Pulmonary Nodules in Computed Tomography Using a Regression Neural Network Approach and its Application to the Lung Image Database Consortium and Image Database Resource Initiative Dataset. TCIA. Saint Louis, MO. (link)
Roth H, Lu L, et al. (2015). A new 2.5D representation for lymph node detection in CT. TCIA. Saint Louis, MO. (link)
Shinagare AB, Vikram R, et al. (2015). Radiogenomics of Clear Cell Renal Cell Carcinoma: Preliminary Findings of The Cancer Genome Atlas-Renal Cell Carcinoma (TCGA-RCC) Research Group. TCIA. Saint Louis, MO. (link)
Vallières M, Freeman CR, et al. (2015). Data from: A radiomics model from joint FDG-PET and MRI texture features for the prediction of lung metastases in soft-tissue sarcomas of the extremities. TCIA. Saint Louis, MO. (link)
Farahani K, Kalpathy-Cramer J, Chenevert TL, et al. Computational Challenges and Collaborative Projects in the NCI Quantitative Imaging Network. Tomography, 2016;2(4):242-9. (link)
Kalpathy-Cramer J, Mamomov A, Zhao B,et al.. Radiomics of Lung Nodules: A Multi-Institutional Study of Robustness and Agreement of Quantitative Imaging Features. Tomography,2016;2(4):430-7. doi: 10.18383/j.tom.2016.00235.
Huang, W., X. Li, et al. (2014). "Variations of dynamic contrast-enhanced magnetic resonance imaging in evaluation of breast cancer therapy response: a multicenter data analysis challenge." Transl Oncol 7(1): 153-166. (link)
Kalpathy-Cramer J, Freymann JB, Kirby JS, et al. Quantitative Imaging Network: Data Sharing and Competitive Algorithm Validation Leveraging The Cancer Imaging Archive Translational Oncology. 2014 Feb;7(1):147-52. doi: 10.1593/tlo.13862. (link)
Lin AY, Du P, Dinning PG, Arkwright JW, Kamp JP, Cheng LK, Bissett IP, O'Grady G. High resolution anatomic correlation of cyclic motor patterns in the human colon: Evidence of a rectosigmoid brake. American Journal of Physiology-Gastrointestinal and Liver Physiology. 2017;312(5):G508-G15. doi: 10.1152/ajpgi.00021.2017.
Gayathri DK, Radhakrishnan R, Rajamani K. Segmentation of colon and removal of opacified fluid for virtual colonoscopy. Pattern Analysis and Applications. 2017:1-15. doi: 10.1007/s10044-017-0614-y
Ryalat MH, Laycock S, Fisher M, editors. Automatic Removal of Mechanical Fixations from CT Imagery with Particle Swarm Optimisation. International Conference on Bioinformatics and Biomedical Engineering; 2017: Springer. DOI: 10.1007/978-3-319-56148-6_37
Farag, A. A., Ali, A., Elshazly, S., & Farag, A. A. (2017). Feature fusion for lung nodule classification. International Journal of Computer Assisted Radiology and Surgery, 1-10. doi:10.1007/s11548-017-1626-1
Mhetre RR, Sache RG. Detection of Lung Cancer Nodule on CT scan Images by using Region Growing Method. International Journal of Current Trends in Engineering & Research. 2016;2(7):215-9. (link)
Setio AAA, Traverso A, de Bel T, Berens MS, Bogaard Cvd, Cerello P, Chen H, Dou Q, Fantacci ME, Geurts B. Validation, comparison, and combination of algorithms for automaticdetection of pulmonary nodules in computed tomography images: the LUNA16 challenge. arXiv preprint arXiv:161208012. 2016:1-16.
Sivakumar, S. and C. Chandrasekar (2015). "A Novel Noise Removal Method for Lung CT SCAN Images Using Statistical Filtering Techniques." International Journal of Algorithms Design and Analysis 1(1).
Magdy, E., N. Zayed, et al. Automatic Classification of Normal and Cancer Lung CT Images using Multi-scale AM-FM Features. Intl Journal of Biomedical Imaging, 2015. (link)
Lassen BC, Jacobs C, et al. Robust Semi-automatic Segmentation of Pulmonary Subsolid Nodules in Chest Computed Tomography Scans. Phys Med Biol (2015) 60(3):1307-1323. (link)
Kumar, D., M. J. Shafiee, et al. Discovery Radiomics for Computed Tomography Cancer Detection. arXiv e-print, 2015. (arXiv link)
Demir, Ö. and A. Yılmaz Çamurcu (2015). "Computer-aided detection of lung nodules using outer surface features." Bio-Medical Materials and Engineering 26(s1): 1213-1222.
Kumar, A., F. Nette, et al. (2014). "A Visual Analytics Approach using the Exploration of Multi-Dimensional Feature Spaces for Content-based Medical Image Retrieval IEEE J Biomed Health Inform (2014) 19(5):1734:1746 (pubmed link)
Raicu DS, Varutbangkul E, Furst JD, Armato SG III: Modeling semantics from image data: Opportunities from LIDC. International Journal of Biomedical Engineering and Technology 3: 83–113, 2010.
Zinovev D, Duo Y, Raicu DS, Furst JD, Armato SG III: Consensus versus disagreement in imaging research: A case study using the LIDC Database. Journal of Digital Imaging 25: 423–436, 2012.
|
|
Please see List of NLST Publications at NIH to browse publications from this Data Collection.
Patil R, Mahadevaiah G, Dekker A. An Approach Toward Automatic Classification of Tumor Histopathology of Non–Small Cell Lung Cancer Based on Radiomic Features. Tomography: a journal for imaging research. 2016;2(4):374-7. (link)
Weis JA, Miga MI, Arlinghaus LR, Li X, Abramson V, Chakravarthy AB, Pendyala P, Yankeelov TE. Predicting the Response of Breast Cancer to Neoadjuvant Therapy Using a Mechanically Coupled Reaction-Diffusion Model. Cancer Res. 2015 Nov 15;75(22):4697-707. doi: 10.1158/0008-5472.CAN-14-2945.
|
Nowaková J, Prílepok M, Snášel V. Medical Image Retrieval Using Vector Quantization and Fuzzy S-tree. Journal of Medical Systems. 2017;41(2):18. (link)
Li, A., C. Li, et al. (2013). Automated Segmentation of Prostate MR Images Using Prior Knowledge Enhanced Random Walker. Digital Image Computing: Techniques and Applications (DICTA), 2013 International Conference on, IEEE.
Qiu, W., J. Yuan, et al. (2014). Prostate segmentation: An efficient convex optimization approach with axial symmetry using 3-D TRUS and MR images. Medical Imaging, IEEE Transactions on 33(4): 947-960.
Xie, Q. and D. Ruan (2014). Low-complexity atlas-based prostate segmentation by combining global, regional, and local metrics. Medical physics 41(4): 041909.
Zhao, T. and D. Ruan (2015). Two-stage fusion set selection in multi-atlas-based image segmentation. Biomedical Imaging (ISBI), 2015 IEEE 12th International Symposium on, IEEE.
Babu, B. S., & Varadarajan, S. (2017). Detection of brain tumour in MRI scan images using Tetrolet Transform and SVM classifier. Indian Journal of Science and Technology, 10. doi:10.17485/ijst/2017/v10i19/113721
Oliveira B, O'Halloran M, Conceicao R, Glavin M, Jones E. Development of Clinically-Informed 3D Tumor Models for Microwave Imaging Applications. IEEE Antennas and Wireless Propagation Letters 2016;15:520-3. DOI: 10.1109/LAWP.2015.2456051
Melouah, A. (2015). Comparison of Automatic Seed Generation Methods for Breast Tumor Detection Using Region Growing Technique. Computer Science and Its Applications, Springer: 119-128.
Desseroit M-C, Visvikis D, Tixier F, Majdoub M, Perdrisot R, Guillevin R, Le Rest CC, Hatt M. Development of a nomogram combining clinical staging with 18F-FDG PET/CT image features in non-small-cell lung cancer stage I–III. European journal of nuclear medicine and molecular imaging. 2016:1-9. DOI: 10.1007/s00259-016-3325-5
|
Wu J, Cui Y, Sun X, Cao G, Li B, Ikeda DM, Kurian AW, Li R. Unsupervised clustering of quantitative image phenotypes reveals breast cancer subtypes with distinct prognoses and molecular pathways. Clinical Cancer Research. 2017:clincanres. 2415.016. (link)
Lavasani, S. N., A. F. Kazerooni, et al. (2015). Discrimination of Benign and Malignant Suspicious BreastTumors Based on Semi-Quantitative DCE-MRI ParametersEmploying Support Vector Machine. Frontiers in Biomedical Technologies 2(2): 397-403.
Anand, S., V. Vinod, et al. Application of Fuzzy c-means and Neural networks to categorize tumor affected breast MR Images. International Journal of Applied Engineering Research 10(64): 2015.
Guo, W., H. Li, et al. (2015). Prediction of clinical phenotypes in invasive breast carcinomas from the integration of radiomics and genomics data. Journal of Medical Imaging 2(4): 041007-041007.
Kim, G. R., Ku, Y. J., Cho, S. G., Kim, S. J., & Min, B. S. (2017). Associations between gene expression profiles of invasive breast cancer and breast imaging reporting and data system MRI lexicon. Annals of Surgical Treatment and Research, 93(1), 18-26. DOI: 10.4174/astr.2017.93.1.18
ParthaSarathi, M., & Ansari, M. A. (2017). Multimodal retrieval framework for brain volumes in 3D MR volumes. Journal of Medical and Biological Engineering, 1-12. doi:10.1007/s40846-017-0287-4
Liu, Y., Xu, X., Yin, L., Zhang, X., Li, L., & Lu, H. (2017). Relationship between glioblastoma heterogeneity and survival time: An MR imaging texture analysis. American Journal of Neuroradiology, 1-7. doi:10.3174/ajnr.A5279.
Beig N, Patel J, Prasanna P, et al. Radiogenomic analysis of hypoxia pathway reveals computerized MRI descriptors predictive of overall survival in Glioblastoma. SPIE Medical Imaging; 2017; 10134:1-10. International Society for Optics and Photonics. doi:10.1117/12.2255694
Lee, J.K., Wang, J., Sa, J.K., et al. Spatiotemporal genomic architecture informs precision oncology in glioblastoma. Nature Genetics.(2017) DOI: 10.1038/ng.3806
Cui Y, Ren S, Tha KK, Wu J, Shirato H, Li R. Volume of high-risk intratumoral subregions at multi-parametric MR imaging predicts overall survival and complements molecular analysis of glioblastoma. European Radiology. 2017:1-10. (link)
Kanas VG, Zacharaki EI, Thomas GA, Zinn PO, Megalooikonomou V, Colen RR. Learning MRI-based classification models for MGMT methylation status prediction in glioblastoma. Computer Methods and Programs in Biomedicine. 2017;140:249-57.(link)
Czarnek N, Clark K, Peters KB, Mazurowski MA. Algorithmic three-dimensional analysis of tumor shape in MRI improves prognosis of survival in glioblastoma: a multi-institutional study. Journal of Neuro-Oncology. 2017:1-8. (link)
Chaddad A, Desrosiers C, Toews M, editors. Radiomic analysis of multi-contrast brain MRI for the prediction of survival in patients with glioblastoma multiforme. Engineering in Medicine and Biology Society (EMBC), 2016 IEEE 38th Annual International Conference; 2016.
Prasanna, P., Patel, J., Partovi, S. et al. Radiomic features from the peritumoral brain parenchyma on treatment-naïve multi-parametric MR imaging predict long versus short-term survival in glioblastoma multiforme: Preliminary findings. Eur Radiol (2016) pp 1–10. DOI:10.1007/s00330-016-4637-3
Mulvey M, Muhyadeen S, Sinha U. Classification of Glioblastoma Multiforme Molecular Subtypes Using Three-Dimensional Multi-Modal MR Imaging Features. Med. Phys. 43, 3373 (2016); (link)
Upadhaya T, Morvan Y, et al. Prognostic value of multimodal MRI tumor features in Glioblastoma multiforme using textural features analysis. In Biomedical Imaging (ISBI), 2015 IEEE 12th International Symposium on, pp. 50-54. IEEE, 2015.
Upadhaya T, Morvan Y, et al. A framework for multimodal imaging-based prognostic model building: Preliminary study on multimodal MRI in Glioblastoma Multiforme. IRBM. 2015 Nov 30;36(6):345-50.
Reza SM, Mays R, Iftekharuddin KM, editors. Multi-fractal detrended texture feature for brain tumor classification. SPIE Medical Imaging; 2015: International Society for Optics and Photonics.
Nabizadeh N, Kubat M. Brain tumors detection and segmentation in MR images: Gabor wavelet vs. statistical features. Computers & Electrical Engineering. 2015.
Natteshan N, Jothi JAA. Automatic Classification of Brain MRI Images Using SVM and Neural Network Classifiers. Advances in Intelligent Informatics: Springer; 2015. p. 19-30. (link)
Zhang J, Barboriak DP, Hobbs H, Mazurowski MA. A fully automatic extraction of magnetic resonance image features in Glioblastoma patients. Medical physics. 2014;41(4):042301.
Wangaryattawanich P, Wang J, Thomas GA, Chaddad A, Zinn PO, Colen RR, editors. Survival analysis of pre-operative GBM patients by using quantitative image features. Control, Decision and Information Technologies (CoDIT), 2014 International Conference on; 2014: IEEE.
Colen RR, Wang J, Singh SK, Gutman DA, Zinn PO. Glioblastoma: Imaging Genomic Mapping Reveals Sex-specific Oncogenic Associations of Cell Death. Radiology. 2014.
Wassal E, Zinn P, Colen R. DIFFUSION AND CONVENTIONAL MR IMAGING GENOMIC BIOMARKER SIGNATURE PREDICTS IDH-1 MUTATION IN GLIOBLASTOMA PATIENTS. Neuro-Oncology. 2014;16(suppl 5):v157-v.
Kwon D, Shinohara RT, Akbari H, Davatzikos C. Combining Generative Models for Multifocal Glioma Segmentation and Registration. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2014: Springer; 2014. p. 763-70.
Wangaryattawanich, P., M. Hatami, et al. "Multicenter imaging outcomes study of The Cancer Genome Atlas glioblastoma patient cohort: imaging predictors of overall and progression-free survival." Neuro-oncology, (2015): nov117 .
Kuo, J. S., K. B. Pointer, et al. (2015). "139 Human Ether-a-Go-Go-Related-1 Gene (hERG) K+ Channel as a Prognostic Marker and Therapeutic Target for Glioblastoma." Neurosurgery 62: 210-211.
Zinn, P. O., M. Hatami, et al. (2015). "138 Diffusion MRI ADC Mapping of Glioblastoma Edema/Tumor Invasion and Associated Gene Signatures." Neurosurgery 62: 210.
Steed, T., J. Treiber, et al. (2015). "Iterative Probabilistic Voxel Labeling: Automated Segmentation for Analysis of The Cancer Imaging Archive Glioblastoma Images." American Journal of Neuroradiology 36(4): 678-685.
Lee, J., S. Narang, et al. (2015). "Associating spatial diversity features of radiologically defined tumor habitats with epidermal growth factor receptor driver status and 12-month survival in glioblastoma: methods and preliminary investigation." Journal of Medical Imaging 2(4): 041006-041006.
Itakura, H., A. S. Achrol, et al. (2015). "Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities." Science Translational Medicine 7(303): 303ra138-303ra138.
Cui, Y., K. K. Tha, et al. (2015). "Prognostic Imaging Biomarkers in Glioblastoma: Development and Independent Validation on the Basis of Multiregion and Quantitative Analysis of MR Images." Radiology: 150358.
Lee, J., S. Narang, et al. (2015). "Spatial Habitat Features Derived from Multiparametric Magnetic Resonance Imaging Data Are Associated with Molecular Subtype and 12-Month Survival Status in Glioblastoma Multiforme." PloS one 10(9): e0136557.
Rios Velazquez E, Meier R, Dunn WD Jr, Alexander B, Wiest R, Bauer S, Gutman DA, Reyes M, Aerts HJ. "Fully automatic GBM segmentation in the TCGA-GBM dataset: Prognosis and correlation with VASARI features." Sci Rep. 2015 Nov 18;5:16822. doi: 10.1038/srep16822.
Collection: TCGA-KIRC