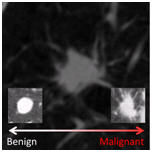
Summary
Update
This challenge was completed in February 2015. An article describing the lessons learned can be found at http://medicalimaging.spiedigitallibrary.org/article.aspx?articleid=2375472. The corresponding scientific manuscript is available at http://dx.doi.org/10.1117/1.JMI.3.4.044506. The full data set, including the answers to the test data, can be found on the SPIE-AAPM-NCI Lung Nodule Classification Challenge Dataset (SPIE-AAPM Lung CT Challenge) page.
As part of the 2015 SPIE Medical Imaging Conference, SPIE – with the support of American Association of Physicists in Medicine (AAPM) and the National Cancer Institute (NCI) – will conduct a “Grand Challenge” on quantitative image analysis methods for the diagnostic classification of malignant and benign lung nodules. The LUNGx Challenge will provide a unique opportunity for participants to compare their algorithms to those of others from academia, industry, and government in a structured, direct way using the same data sets.
LUNGx Challenge Format
Release of the calibration dataset (with truth): November 21, 2014
- 10 contrast-enhanced CT scans will be available as a calibration dataset. This dataset is representative of the technical properties (scanner type, acquisition parameters, file format) of the test dataset. Participants should not necessarily consider the lung nodules present in the calibration cases to be representative of the difficulty level expected in the test set.
- The calibration set contains five CT scans with malignant nodules and five CT scans with benign nodules. The organizers have selected a single nodule per CT scan for analysis.
- The location of each nodule is specified in an associated Excel file that includes case name, the coordinates of the approximate nodule centroid, and the ‘truth’ label (malignant or benign).
- Participants are encouraged to calibrate their algorithms by downloading the calibration dataset through the TCIA.
Dataset Download
- Image data - Save/open this file to initiate our Java Web Start download manager to begin your download
- Nodule locations and 'truth' labels
Disclaimer: Anyone wishing to use the downloaded images for presentation or publication purposes outside of the LUNGx Challenge should acknowledge the SPIE, the NCI, the AAPM, and The University of Chicago:
“Data used in this research were obtained from The Cancer Imaging Archive (TCIA) sponsored by the SPIE, NCI/NIH, AAPM and The University of Chicago.”
Please also cite both http://dx.doi.org/10.7937/K9/TCIA.2015.UZLSU3FL and http://medicalimaging.spiedigitallibrary.org/article.aspx?articleid=2375472
Release of the test dataset (without truth): January 9, 2015
The 10 calibration cases released in November were made available specifically so that participants could become familiar with the image acquisition parameters and DICOM files of the one institution that is supplying the clinical cases for the LUNGx Challenge (The University of Chicago). The calibration cases were intended to orient participants to relevant technical issues surrounding the compatibility of the Challenge cases with participants’ algorithms and were not intended for algorithm development or training.
The 60 CT scans that comprise the LUNGx Challenge test set are now available. Most of these cases are contrast-enhanced scans. The location of each nodule for analysis is specified in the accompanying Excel spreadsheet that includes case name, the number of nodules for analysis in the scan (either 1 or 2), and the coordinates of each nodule’s approximate centroid. Important reference and explanatory information appears at the bottom of this spreadsheet. The nodules contained in the test set cases have been determined by a radiologist to be either primary lung cancer or benign based on pathologic assessment and/or follow-up imaging examinations.
A Google Group mailing list has been established as the primary mechanism through which news, updates, and answers to questions will be communicated by the organizers of the Challenge. It is essential that all participants sign up to receive these emails at: https://groups.google.com/forum/#!forum/lungx-challenge. This URL may also be used to view the archive of prior announcements. Questions about the Challenge should be directed to Sam Armato (s-armato@uchicago.edu) or Karen Drukker (kdrukker@uchicago.edu); please note that answers to any questions will be distributed only through the Google Group mailing list.
No more than two sets of results obtained from distinct methods may be submitted by a single group along with a description of the methods used. By submitting results for the LUNGx Challenge, participants agree that their results (along with a very brief description of their method) may be presented as part of publications prepared about the LUNGx Challenge by the organizers.
Dataset Download
- Test Image Data - Save/open this file to initiate our Java Web Start download manager to begin your download
- Test Nodule locations
Disclaimer: Anyone wishing to use the downloaded images for presentation or publication purposes outside of the LUNGx Challenge should acknowledge the SPIE, the NCI, the AAPM, and The University of Chicago:
“Data used in this research were obtained from The Cancer Imaging Archive (TCIA) sponsored by the SPIE, NCI/NIH, AAPM and The University of Chicago.”
Please also cite both http://dx.doi.org/10.7937/K9/TCIA.2015.UZLSU3FL and http://medicalimaging.spiedigitallibrary.org/article.aspx?articleid=2375472
Deadline for participants to submit test set classification results: February 6, 2015
- Participants should submit their test set classification results by e-mail to Karen Drukker (kdrukker@uchicago.edu). In the e-mail
- The subject line it should read: 'LUNGx Challenge' .
- The text should include (1) a clear identification of the participant(s) by name and affiliation (research group/institution/company), and (2) a brief description of the employed CADx method (with citations to the literature if applicable).
- Results should be submitted as an attachment (for specific formatting specifications please see detailed instructions below).
- An acknowledgment will be sent within 2 business days of receipt of results. In case no confirmation of receipt is received please contact customerservice@spie.org.
Output format for the LUNGx Challenge test set classification results:
In order to facilitate performance assessment of your method for the test cases in the LUNGx Challenge (and to compare your performance to that of the methods of other participants), please follow these guidelines:
- Using your method, calculate a ‘score’ for each case (nodule) assessing the level of suspicion that this case is malignant (or benign).
- Note that the performance of your method will be assessed using Receiver Operator Characteristic (ROC) analysis with the area under the ROC curve as a figure of merit. Hence, a continuous scale of your output ‘score’ is preferred (for example, continuous from 0 to 1). A binary ‘score’ (such as 0=benign and 1=malignant with nothing in between) is NOT acceptable. Categorical output is not desirable but may be acceptable if a sufficiently large number of categories in a logical order is used.
- Tabulate your ‘scores’ for all cases in a file containing 2 columns with
- The first column being the case name, the second column being the nodule number, and the third column your ‘score’. Example:
- LUNGx-CT001 1 0.034
- LUNGx-CT002 1 0.001
- LUNGx-CT002 2 0.95
- LUNGx-CT003 1 0.851
- The preferred file format is Excel, but CSV and OpenOffice (.sxc) also are acceptable.
- The first column being the case name, the second column being the nodule number, and the third column your ‘score’. Example:
- Provide a note at the top of your tabulated list whether, for your method, high or low values for the ‘score’ are indicative of malignancy. For example, if your ‘score’ is a probability of malignancy then a higher ‘score’ is expected for actually malignant cases, and hence you would note that ‘high’ values indicate malignancy.
Organizers and Major Contributors:
- Samuel G. Armato, University of Chicago (s-armato@uchicago.edu)
- Lubomir Hadjiiski, University of Michigan Health System (lhadjisk@umich.edu)
- Georgia Tourassi, Oak Ridge National Lab. (tourassig@ornl.gov)
- Karen Drukker, University of Chicago (kdrukker@uchicago.edu)
- Maryellen Giger, University of Chicago (m-giger@uchicago.edu)
Feng Li, University of Chicago (feng@uchicago.edu)
- George Redmond, NIH/NCI (gr34m@nih.gov)
- Laurence Clarke, NIH/NCI (lclarke@mail.nih.gov)
- Keyvan Farahani, NIH/NCI (farahank@mail.nih.gov)
- Justin Kirby, Leidos Biomedical, Inc (kirbyju@mail.nih.gov)
- Angela Keyser, AAPM (akeyser@aapm.org)
- Diane Cline, SPIE (diane@spie.org)
- Sandy Hoelterhoff, SPIE (sandyh@spie.org)