- Created by Sean Berryman, last modified by Jeff Tobler on Apr 12, 2024
Summary
Redirection Notice
This page will redirect to https://www.cancerimagingarchive.net/collection/pancreas-ct/ in about 5 seconds.
A medical student manually performed slice-by-slice segmentations of the pancreas as ground-truth and these were verified/modified by an experienced radiologist.
Data Access
| Data Type | Download all or Query/Filter | License |
|---|---|---|
| Images (DICOM, 9.3 GB) | (Download requires NBIA Data Retriever) | |
| Manual Annotations (zip, 975 kB) |
Click the Versions tab for more info about data releases.
Additional Resources for this Dataset
The NCI Cancer Research Data Commons (CRDC) provides access to additional data and a cloud-based data science infrastructure that connects data sets with analytics tools to allow users to share, integrate, analyze, and visualize cancer research data.
- Imaging Data Commons (IDC) (Imaging Data)
Detailed Description
Collection Statistics | |
|---|---|
Modalities | CT |
Number of Participants | 80 |
Number of Studies | 80 |
Number of Series | 80 |
Number of Images | 18,942 |
| Image Size (GB) | 9.3 |
Data Example
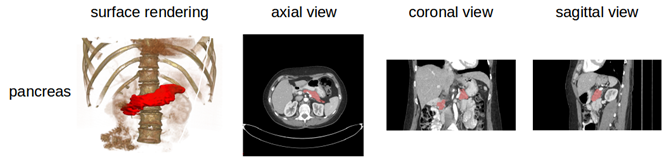
Note
The DICOM files were created from anonymized volumetric images (Analyze and NifTI) using this from ITK: http://www.itk.org/Doxygen/html/Examples_2IO_2ImageReadDicomSeriesWrite_8cxx-example.html .
Citations & Data Usage Policy
Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution should include references to the following citations:
Data Citation
Roth, H., Farag, A., Turkbey, E. B., Lu, L., Liu, J., & Summers, R. M. (2016). Data From Pancreas-CT (Version 2) [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/K9/TCIA.2016.tNB1kqBU
Publication Citation
Roth HR, Lu L, Farag A, Shin H-C, Liu J, Turkbey EB, Summers RM. DeepOrgan: Multi-level Deep Convolutional Networks for Automated Pancreas Segmentation. N. Navab et al. (Eds.): MICCAI 2015, Part I, LNCS 9349, pp. 556–564, 2015. (arXiv link) https://doi.org/10.1007/978-3-319-24553-9_68
TCIA Citation
Clark K, Vendt B, Smith K, Freymann J, Kirby J, Koppel P, Moore S, Phillips S, Maffitt D, Pringle M, Tarbox L, Prior F. The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository, Journal of Digital Imaging, Volume 26, Number 6, December, 2013, pp 1045-1057. DOI: https://doi.org/10.1007/s10278-013-9622-7
Other Publications Using This Data
TCIA maintains a list of publications that leverage TCIA data. If you have a manuscript you'd like to add please contact the TCIA Helpdesk. Below is a list of such publications using this Collection:
- Gibson, E., Giganti, F., Hu, Y., Bonmati, E., Bandula, S., Gurusamy, K., . . . Barratt, D. C. (2017). Towards Image-Guided Pancreas and Biliary Endoscopy: Automatic Multi-organ Segmentation on Abdominal CT with Dense Dilated Networks. Paper presented at the International Conference on Medical Image Computing and Computer-Assisted Intervention.
- Greenspan, H., van Ginneken, B., & Summers, R. M. (2016). Guest Editorial Deep Learning in Medical Imaging: Overview and Future Promise of an Exciting New Technique. IEEE Transactions on Medical Imaging, 35(5), 1153-1159. doi:10.1109/TMI.2016.2553401
- Shi, H., Lu, L., Yin, M., Zhong, C., & Yang, F. (2023). Joint few-shot registration and segmentation self-training of 3D medical images. Biomedical Signal Processing and Control, 80. doi:https://doi.org/10.1016/j.bspc.2022.104294
Version 2 (Current): Updated 2020/09/10
| Data Type | Download all or Query/Filter |
|---|---|
| Images (DICOM, 9.3 GB) | (Download requires the NBIA Data Retriever) |
| Manual Annotations |
Note: Previously posted cases #25 and #70 were found to be from the same scan as case #2, just cropped slightly differently, and were removed from this version of the dataset.
Version 1 : Updated 2015/12/29
- No labels