Summary
| Excerpt | ||||||||
|---|---|---|---|---|---|---|---|---|
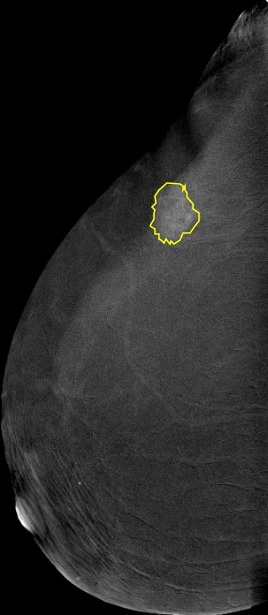
reducing the workload of radiologists and helping them provide a more accurate diagnosis. However, fully annotated and large-sized datasets are required. In the past couple of years, a few public mammography datasets were released. These datasets contain digital mammography images only, and none include CESM images.This dataset is a collection of 2,006 high-resolution Contrast-enhanced spectral mammography (CESM) images with annotations and medical reports. Acquisition protocol:CESM is done using the standard DMdigital mammography equipment but, with additional software that performs dual-energy image acquisition. TwoTwo minutes after intravenously injecting the patient with non-ionic low-osmolar iodinated contrast material (dose: 1.5 mL/kg), craniocaudalcraniocaudal (CC) and mediolateral oblique (MLO) views are obtained. Each view comprises two exposures, one with low energylow energy (peak kilo-voltage values ranging from 26 to 31kVp) and one with high energy (45 to 49 kVp). Low and high-energy images are then recombined and subtracted through appropriate image processing to suppress the background breast parenchyma. A complete examinationexamination is carried out in about 5-6 minutes. Image preprocessing:The images were converted from DICOM to JPEG using RadiAnt with best 100% image quality (lossless). They have an average of 2355 x 1315 pixels. Supporting data:Full medical reports are also provided for each case (DOCX) along with manual segmentation annotation for the abnormal findings in each image (CSV file). Each image with its corresponding manual annotation (breast composition, mass shape, mass margin, mass density, architectural distortion, asymmetries, calcification type, calcification distribution, mass enhancement pattern, non-mass enhancement pattern, non-mass enhancement distribution, and overall BIRADS assessment) is compiled into 1 Excel file. https://www.robots.ox.ac.uk/~vgg/software/via/via.html was used for the segmentation annotation. It can be used to show the annotations on the images by clicking on Annotation--> import annotations (from csv), and then proceeding to upload any image to view the annotations drawn over it. Moreover, a helper repository is created to help with pre-processing, model training, model evaluation, and segmentation annotation loading: https://github.com/omar-mohamed/CDD-CESM-Dataset Regarding the tabs on the annotations Excel file, these are commonly used radiological descriptors as defined by the American College of Radiology 2013 lexicon. |
Acknowledgements
We would like to acknowledge the individuals and institutions that have provided data for this collection:
- National Cancer Institute, Cairo University, Cairo, Egypt : Special thanks to Dr. Rana Khaled, M.Sc, Prof. Maha Helal, MD, Prof. Omnia Mokhtar, MD and Dr. Hebatalla El Kassas, MD from the Department of Radiology.
- Faculty of Computers and Artificial Intelligence, Cairo University, Cairo, Egypt – Special thanks to Omar Alfarghaly, Prof. Abeer Elkorany, and Prof. Aly Fahmy from the Department of Computer Science
Hospital/Institution Name city, state, country - Special thanks to First Last Names, degree PhD, MD, etc from the Department of xxxxxx, Additional Names from same location.
- Continue with any names from additional submitting sites if collection consists of more that one.
| Localtab Group | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|