Versions Compared
Key
- This line was added.
- This line was removed.
- Formatting was changed.
Summary
| Excerpt |
|---|
This collection is a retrospective set of prostate MR studies. All studies included T2-weighted (T2W), proton density-weighted (PD-W), dynamic contrast enhanced (DCE), and diffusion-weighted (DW) imaging. The images were acquired on two different types of Siemens 3T MR scanners, the MAGNETOM Trio and Skyra. T2-weighted images were acquired using a turbo spin echo sequence and had a resolution of around 0.5 mm in plane and a slice thickness of 3.6 mm. The DCE time series was acquired using a 3-D turbo flash gradient echo sequence with a resolution of around 1.5 mm in-plane, a slice thickness of 4 mm and a temporal resolution of 3.5 s. The proton density weighted image was acquired prior to the DCE time series using the same sequence with different echo and repetition times and a different flip angle. Finally, the DWI series were acquired with a single-shot echo planar imaging sequence with a resolution of 2 mm in-plane and 3.6 mm slice thickness and with diffusion-encoding gradients in three directions. Three b-values were acquired (50, 400, and 800), and subsequently, the ADC map was calculated by the scanner software. All images were acquired without an endorectal coil. |
Accessing the PROSTATEx Challenge Data Sets
The PROSTATEx Challenge ("SPIE-AAPM-NCI Prostate MR Classification Challenge”) focused on quantitative image analysis methods for the diagnostic classification of clinically significant prostate cancers and was held in conjunction with the 2017 SPIE Medical Imaging Symposium (see http://www.spie.org/PROSTATEx/). PROSTATEx ran from November 21, 2016 to January 15, 2017, though a "live" version has also been established at https://prostatex.grand-challenge.org which serves as an ongoing way for researchers to benchmark their performance for this task.
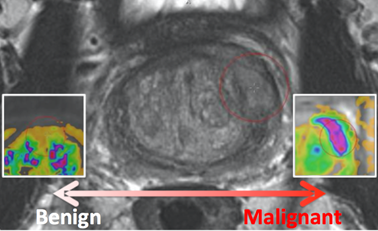
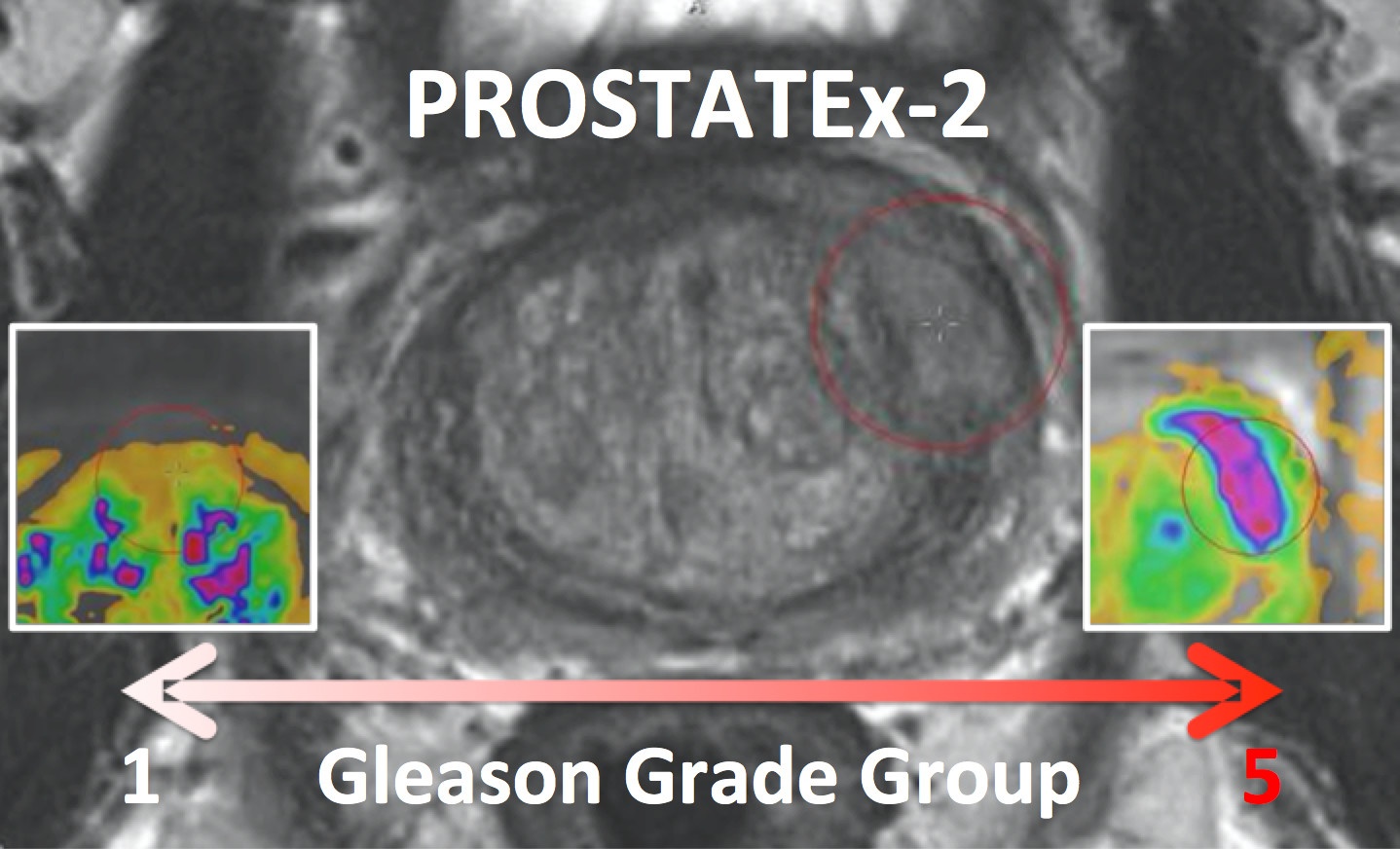
The PROSTATEx-2 Challenge ("SPIE-AAPM-NCI Prostate MR Gleason Grade Group Challenge") ran from May 15, 2017 to June 23, 2017 and was focused on the development of quantitative multi-parametric MRI biomarkers for the determination of Gleason Grade Group in prostate cancer. It was held in conjunction with the 2017 AAPM Annual Meeting (see http://www.aapm.org/GrandChallenge/PROSTATEx-2/default.asp).
Supplemental data and instructions specific to both challenges can be found on the Detailed Description tab below.
Acknowledgements
The prostate MR imaging was performed at the Radboud University Medical Centre (Radboudumc) in the Prostate MR Reference Center under supervision of prof. Dr. Barentsz. The Radboudumc is located in Nijmegen, The Netherlands. The dataset was collected and curated for research in computer aided diagnosis of prostate MR under supervision of Dr. Huisman, Radboudumc.
| Localtab Group | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|