Summary
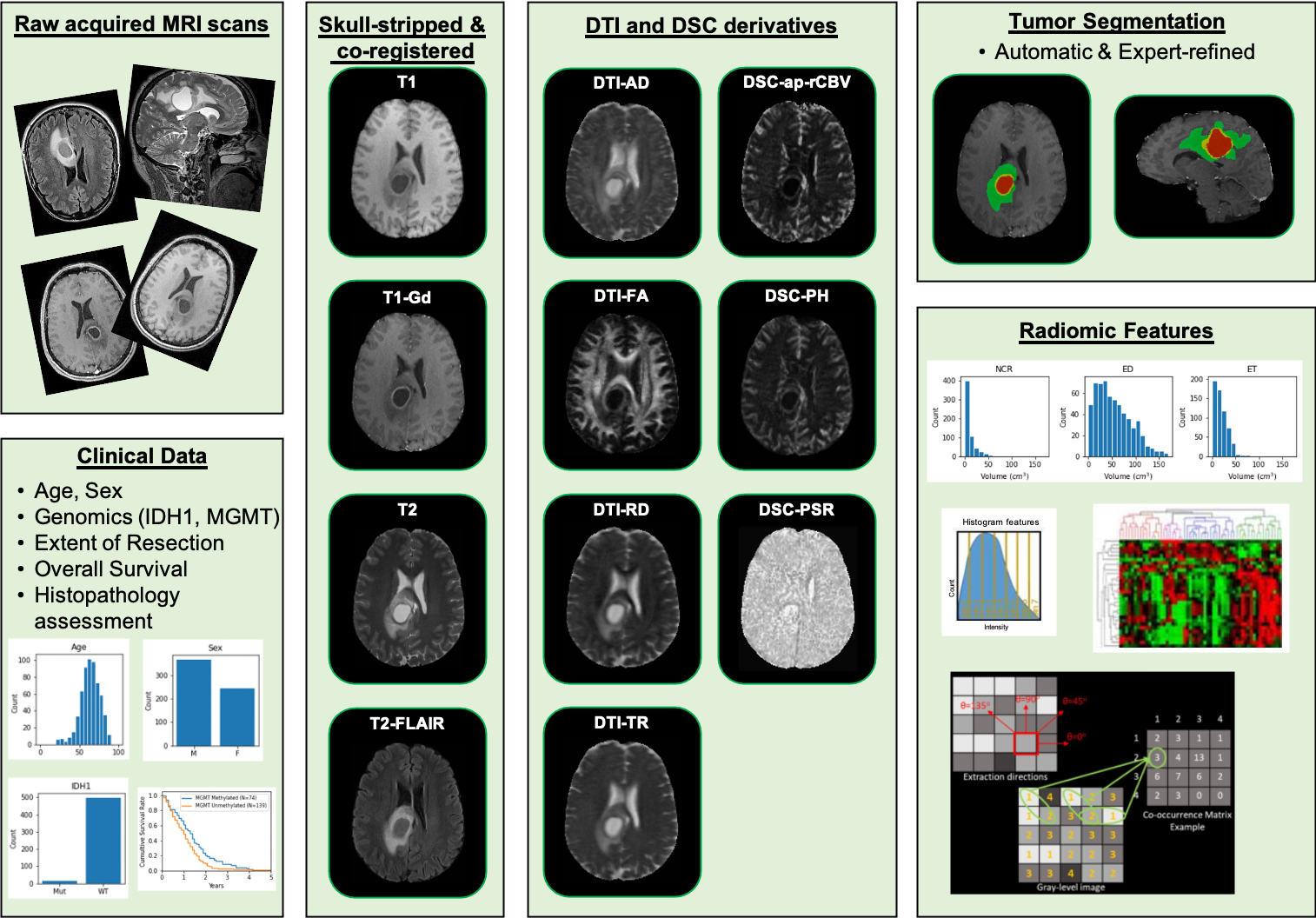
This collection comprises multi-parametric magnetic resonance imaging (mpMRI) scans for de novo Glioblastoma (GBM) patients from the University of Pennsylvania Health System, coupled with patient demographics, clinical outcome (e.g., overall survival, genomic information, tumor progression), as well as computer-aided and manually-corrected segmentation labels of multiple histologically distinct tumor sub-regions, computer-aided and manually-corrected segmentations of the whole brain, a rich panel of radiomic features along with their corresponding co-registered mpMRI volumes in NIfTI format. Scans were initially skull-stripped and co-registered, before their tumor segmentation labels were produced by an automated computational method. These segmentation labels were revised and any label misclassifications were manually corrected/approved by expert board-certified neuroradiologists. The final labels were used to extract a rich panel of imaging features, including intensity, volumetric, morphologic, histogram-based and textural parameters. The segmentation labels enable quantitative computational and clinical studies without the need to repeat manual annotations whilst allowing for comparison across studies. They can also serve as a set of manually-annotated gold standard labels for performance evaluation in computational challenges. The provided panel of radiomic features may facilitate research integrative of the molecular characterization offered, and hence allow associations with molecular markers (radiogenomic biomarker research), clinical outcomes, treatment responses and other endpoints, by researchers without sufficient computational background to extract such features. Additional data accompanying the UPENN-GBM data collection include H&E-stained digitized tissue sections from resected tumor specimens of matched de novo and recurrent cases for a few of the patients in this collection.
Acknowledgements
Reported research was partly supported by the National Cancer Institute (NCI), the National Institute of Neurological Disorders and Stroke (NINDS), and the National Center for Advancing Translational Sciences (NCATS) of the National Institutes of Health (NIH) under award numbers NINDS:R01NS042645, NCI:U24CA189523, NCI:U01CA242871, NCATS:UL1TR001878, and by the Institute for Translational Medicine and Therapeutics (ITMAT) of the University of Pennsylvania. The content of this publication is solely the responsibility of the authors and does not represent the official views of the NIH, or the ITMAT of the UPenn.
Data Access
| Data Type | Download all or Query/Filter | License |
|---|---|---|
Images (DICOM, 139.4 GB) | (Download requires the NBIA Data Retriever) | |
Images (NIfTI, 69 GB) | (Download and apply the IBM-Aspera-Connect plugin to your browser to retrieve this faspex package) | |
Histopathology Images (NDPI, 149 GB) | (Download and apply the IBM-Aspera-Connect plugin to your browser to retrieve this faspex package) | |
| Clinical Data (CSV, 51 kB) | ||
| Histopathology to Radiology Filename Mapping (CSV, 2 kB) | ||
| Image acquisition parameters (CSV, 195 kB) | ||
| Data availability per subject (CSV, 126 kB) | ||
| CaPTk radiomic features list (CSV, 6 kB) | ||
| CaPTk radiomic feature parameter file (CSV, 4 kB) | ||
| Radiomic Data (ZIP,15.37 MB) |
Click the Versions tab for more info about data releases.
Please contact help@cancerimagingarchive.net with any questions regarding usage.
Additional Resources for this Dataset
The NCI Cancer Research Data Commons (CRDC) provides access to additional data and a cloud-based data science infrastructure that connects data sets with analytics tools to allow users to share, integrate, analyze, and visualize cancer research data.
- Imaging Data Commons (IDC) (Imaging Data)
Detailed Description
Image Statistics | ||
|---|---|---|
Modalities | MR | Pathology |
Number of Patients | 630 | 34 |
Number of Studies | 3,301 | N/A |
Number of Series | 3,680 | N/A |
Number of Images | 828,234 | 71 |
| Images Size (GB) | 139.4 | 149 |
Note from the submitting group: The NIfTI images are all registered to a common atlas (SRI) using a uniform preprocessing and the segmentation are aligned with them. Therefore the NIfTI images will not align with the DICOM images, by design. If you load the NIfTI images (like T1/T2) and their related segmentation, these will line up.
Citations & Data Usage Policy
Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution should include references to the following citations:
Data Citation
Bakas, S., Sako, C., Akbari, H., Bilello, M., Sotiras, A., Shukla, G., Rudie, J. D., Flores Santamaria, N., Fathi Kazerooni, A., Pati, S., Rathore, S., Mamourian, E., Ha, S. M., Parker, W., Doshi, J., Baid, U., Bergman, M., Binder, Z. A., Verma, R., … Davatzikos, C. (2021). Multi-parametric magnetic resonance imaging (mpMRI) scans for de novo Glioblastoma (GBM) patients from the University of Pennsylvania Health System (UPENN-GBM) (Version 2) [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/TCIA.709X-DN49
Publication Citation
Bakas, S., Sako, C., Akbari, H., Bilello, M., Sotiras, A., Shukla, G., Rudie, J. D., Flores Santamaria, N., Fathi Kazerooni, A., Pati, S., Rathore, S., Mamourian, E., Ha, S. M., Parker, W., Doshi, J., Baid, U., Bergman, M., Binder, Z. A., Verma, R., Lustig, R., Desai, A. S., Bagley, S. J., Mourelatos, Z., Morrissette, J., Watt, C. D., Brem, S., Wolf, R. L., Melhem, E. R., Nasrallah, M. P., Mohan, S., O’Rourke, D. M., Davatzikos, C. (2022). The University of Pennsylvania glioblastoma (UPenn-GBM) cohort: advanced MRI, clinical, genomics, & radiomics. In Scientific Data (Vol. 9, Issue 1). https://doi.org/10.1038/s41597-022-01560-7
TCIA Citation
Clark, K., Vendt, B., Smith, K., Freymann, J., Kirby, J., Koppel, P., Moore, S., Phillips, S., Maffitt, D., Pringle, M., Tarbox, L., & Prior, F. (2013). The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository. Journal of Digital Imaging, 26(6), 1045–1057. https://doi.org/10.1007/s10278-013-9622-7
Other Publications Using This Data
TCIA maintains a list of publications which leverage TCIA data. If you have a manuscript you'd like to add please contact the TCIA Helpdesk.
Version 2 (Current): 2022/10/24
Note: 2023/12/5: updated clinical data file (v1.2) to include more information about censor/survival.
| Data Type | Download all or Query/Filter | License |
|---|---|---|
Images (DICOM, 139.4 GB) | (Download requires the NBIA Data Retriever) | |
Images (NIfTI, 69 GB) | (Download and apply the IBM-Aspera-Connect plugin to your browser to retrieve this faspex package) | |
Histopathology Images (NDPI, 149 GB) | (Download and apply the IBM-Aspera-Connect plugin to your browser to retrieve this faspex package) | |
| Clinical Data (CSV, 51 kB) | ||
| Histopathology to Radiology Filename Mapping (CSV, 2kB) | ||
| Image acquisition parameters (CSV, 195 kB) | ||
| Data availability per subject (CSV, 126 kB) | ||
| CaPTk radiomic features list (CSV, 6 kB) | ||
| CaPTk radiomic feature parameter file (CSV, 4 kB) | ||
| Radiomic Data (ZIP,15.37 MB) |
changes: Histopathology NDPI slides added to collection. CSV file for mapping Radiology subject IDs to Histopathology patient and image IDs where available (note: not all Radiology data has associated pathology data and vice versa).
Version 1: 2022/06/21
| Data Type | Download all or Query/Filter | License |
|---|---|---|
Images (DICOM, 139.4 GB) | (Download requires the NBIA Data Retriever) | |
Images (NIfTI, 69 GB) | (Download and apply the IBM-Aspera-Connect plugin to your browser to retrieve this faspex package) | |
| Clinical Data (CSV, 51 kB) | ||
| Image acquisition parameters (CSV, 195 kB) | ||
| Data availability per subject (CSV, 126 kB) | ||
| CaPTk radiomic features list (CSV, 6 kB) | ||
| CaPTk radiomic feature parameter file (CSV, 4 kB) | ||
| Radiomic Data (ZIP,15.37 MB) | ||
| CaPTk radiomic feature parameter (CSV, 5KB) |