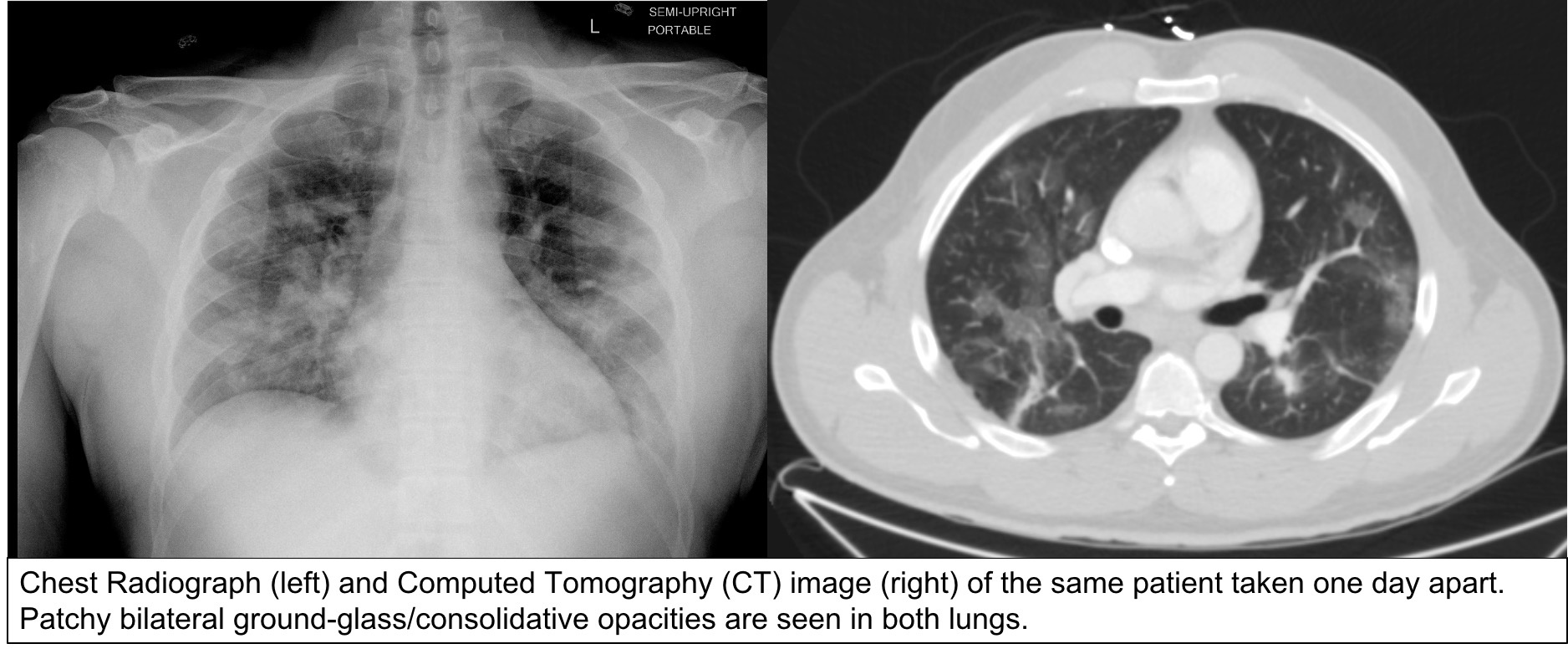
Summary
TCIA COVID-19 Datasets
Additional datasets and information about TCIA efforts to support COVID-19 research can be found here.
Acknowledgements
We would like to acknowledge the individuals and institutions that have provided data for this collection:
The University of Arkansas for Medical Sciences (UAMS) Translational Research Institute, Department of Radiology, Department of Biomedical Informatics and Department of Surgery, Little Rock, Arkansas, USA.
Data Access
| Data Type | Download all or Query/Filter | License |
|---|---|---|
Images (DICOM, 19.0 GB) | (Download requires the NBIA Data Retriever) | |
| Clinical data (CSV, 46 kB) |
Click the Versions tab for more info about data releases.
Please contact help@cancerimagingarchive.net with any questions regarding usage.
Additional Resources for this Dataset
The following external resources have been made available by the data submitters. These are not hosted or supported by TCIA, but may be useful to researchers utilizing this collection.
- Viral Genomes Genbank repository (accession no. MT766907: USA/AR-UAMS001/2020)
- Viral Genomes Genbank repository (accession no. MT766908: USA/AR-UAMS002/2020)
The NCI Cancer Research Data Commons (CRDC) provides access to additional data and a cloud-based data science infrastructure that connects data sets with analytics tools to allow users to share, integrate, analyze, and visualize cancer research data.
- Imaging Data Commons (IDC) (Imaging Data)
Detailed Description
Image Statistics | |
|---|---|
Modalities | CT, CR, DX |
Number of Patients | 105 |
Number of Studies | 256 |
Number of Series | 461 |
Number of Images | 31,935 |
| Images Size (GB) | 19.0 |
Citations & Data Usage Policy
Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution should include references to the following citations:
Data Citation
Desai, S., Baghal, A., Wongsurawat, T., Al-Shukri, S., Gates, K., Farmer, P., Rutherford, M., Blake, G.D., Nolan, T., Powell, T., Sexton, K., Bennett, W., Prior, F. (2020). Data from Chest Imaging with Clinical and Genomic Correlates Representing a Rural COVID-19 Positive Population [Data set]. The Cancer Imaging Archive. DOI: https://doi.org/10.7937/tcia.2020.py71-5978.
Acknowledgement
This project has been funded in whole or in part with federal funds from the National Center for Advancing Translational Sciences UL1 TR003107 and the National Cancer Institute, Contract No. 75N91019D00024, Subcontract 20X023F.
Publication Citation
Desai, S., Baghal, A., Wongsurawat, T., Jenjaroenpun, P., Powell, T., Al-Shukri, S., Gates, K., Farmer, P., Rutherford, M., Blake, G., Nolan, T., Sexton, K., Bennett, W., Smith, K., Syed, S., Prior, F. (2020). Chest imaging representing a COVID-19 positive rural U.S. population. Scientific Data. 2020;7(1):414. doi: https://doi.org/10.1038/s41597-020-00741-6.
Publication Citation
Jenjaroenpun, P., Wanchai, V., OnoMoore, K.D., Laudadio, J., James, L.P., Adams, S.H., Prior, F., Nookaew, I., Ussery, D.W., Wongsurawat, T. (2020). Two SARS-CoV-2 genome sequences of isolates from rural U.S. patients harboring the D614G mutation, obtained using Nanopore sequencing. Microbiology Resource Announcements, 2020. DOI: https://doi.org/10.1128/MRA.01109-20.
TCIA Citation
Clark, K., Vendt, B., Smith, K., Freymann, J., Kirby, J., Koppel, P., Moore, S., Phillips, S., Maffitt, D., Pringle, M., Tarbox, L., & Prior, F. (2013). The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository. In Journal of Digital Imaging (Vol. 26, Issue 6, pp. 1045–1057). Springer Science and Business Media LLC. https://doi.org/10.1007/s10278-013-9622-7 PMCID: PMC3824915
Other Publications Using This Data
TCIA maintains a list of publications which leverage our data. If you have a manuscript you'd like to add please contact TCIA's Helpdesk.
Version 1 (Current): Updated 2020/07/13
| Data Type | Download all or Query/Filter |
|---|---|
Images (DICOM, 19.0 GB) | (Requires NBIA Data Retriever.) |
| Clinical Data (CSV) |
Additional Resources for this Dataset
- Viral Genomes Genbank repository (accession no. MT766907: USA/AR-UAMS001/2020)
- Viral Genomes Genbank repository (accession no. MT766908: USA/AR-UAMS002/2020)