Summary
I-SPY 2 (Investigation of Serial studies to Predict Your Therapeutic Response with Imaging And moLecular analysis 2) is an ongoing, multi-center trial (NCT01042379) designed to quickly evaluate the efficacy of new agents for breast cancer in neoadjuvant chemotherapy (NAC) setting. Women aged ≥18 years diagnosed with locally advanced breast cancer (tumor size ≥2.5 cm) without distant metastasis are eligible to enroll in the trial. Patients with breast cancer at high-risk for recurrence are adaptively-randomized to either control arm (standard NAC) or one of several concurrent experimental drug arms. I-SPY 2 utilizes change in tumor volume by DCE-MRI at serial time-points during NAC to adjust the randomization schema as the trial proceeds, preferentially assigning patients to receive agents showing an increasing likelihood of efficacy against their breast cancer subtype. I-SPY 2 opened in Spring 2010 and is ongoing.
This collection includes DCE MRI data (original acquired images and derived parametric maps) for I-SPY 2 patients adaptively randomized between 2010 and 2016, along with histopathologic outcome data. Breast MRI data in this collection was acquired prospectively at over 22 clinical centers using a standardized image acquisition protocol. Patients underwent 4 MRI exams before and during NAC and over 95% of the DCE imaging data met acceptance criteria for analysis of functional tumor volume (FTV), a demonstrated quantitative imaging marker of breast cancer response to NAC. This is a comprehensive, highly curated imaging data set with histopathologic outcome that can be used to develop, test and compare imaging metrics and prediction models for breast cancer response to treatment. Original but uncurated T2weighted MRI images are also included for most studies.
719 patients are included in this collection. In combination with 266 patients from the ACRIN-6698/ISPY2 TCIA collection these 985 patients comprise "I-SPY2 Imaging Cohort 1", the 1st subgroup of publicly released imaging data from the I-SPY 2 TRIAL. All subjects in the I-SPY2 Imaging Cohort 1 were enrolled in I-SPY 2 from 2010 to 2016 and randomized to one of nine completed experimental drug arms or control arm. In order to obtain the MRI data for the entire I-SPY2 Imaging Cohort 1 please download using the manifest file provided in the Data Access table below as this will include the patients from both collections. Downloading this collection alone will result in an incomplete cohort of 719 patients and any results from that patient set will not be comparable to published results from the trial.
In addition, a retrospective study using a 384 patient subset (median age: 49 y/o) was performed to test if prediction models combining multiple MRI features outperform models with single features. Four features were quantitatively calculated in each MRI exam: functional tumor volume, longest diameter, sphericity, and contralateral background parenchymal enhancement. Logistic regression analysis was used to study the relationship between MRI variables and pathologic complete response (pCR). Predictive performance was estimated using the area under the receiver operating characteristic curve (AUC). The cohort was stratified by hormone receptor (HR) and human epidermal growth factor receptor 2 (HER2) status (positive or negative). Results showed analysis with combined features achieved higher AUCs than analysis with any feature alone. These results are further described in the associated publication and in the multi-feature MRI spreadsheet included with this Collection.
I-SPY2 Imaging Biomarker Group
We are establishing the I-SPY 2 Imaging Biomarker Subgroup to provide a forum where imaging scientists and radiologists can explore, discuss, and strategize on the use of I-SPY 2 MRI data available through TCIA. Please access https://forms.office.com/r/u0xzwB9kbU to be considered to join our bimonthly Zoom meeting invite list.
Acknowledgements
The I-SPY2 Breast MRI Collection is supported by NIH grants U01 CA225427, R01 CA132870 and P01 CA210961. The I-SPY 2 TRIAL is supported by the Quantum Leap Healthcare Collaborative. The authors gratefully acknowledge and thank the patients who have volunteered to participate in the I-SPY2 TRIAL, as well as the extensive network of investigators, patient advocates and study coordinators. We especially thank the site radiology teams that have contributed substantially to the value of this archive.
Data Access
| Data Type | Download all or Query/Filter | License |
|---|---|---|
Images (DICOM, 1.6 TB) 719 patients | (Download requires the NBIA Data Retriever) | |
Images (DICOM, 2.4 TB) I-SPY2 Imaging Cohort 1 dataset, 985 patients (719 I-SPY2 patients plus 266 patients from the ACRIN-6698/ISPY2 TCIA collection) | (Download requires the NBIA Data Retriever) | |
Clinical data (CSV, 57 kB) I-SPY2 Imaging Cohort 1 dataset, 985 patients (719 I-SPY2 patients plus 266 patients from the ACRIN-6698/ISPY2 TCIA collection) | ||
Multi-feature MRI NACT Data (.xlsx, 130 kB) “Predicting breast cancer response to neoadjuvant treatment using multi-feature MRI: results from the I-SPY 2 TRIAL” cohort, 384 patients |
Click the Versions tab for more info about data releases.
Please contact help@cancerimagingarchive.net with any questions regarding usage.
Additional Resources for this Dataset
The NCI Cancer Research Data Commons (CRDC) provides access to additional data and a cloud-based data science infrastructure that connects data sets with analytics tools to allow users to share, integrate, analyze, and visualize cancer research data.
- Imaging Data Commons (IDC) (Imaging Data)
Detailed Description
Image Statistics | |
|---|---|
Modalities | MR,SEG |
Number of Patients | 719 |
Number of Studies | 2688 |
Number of Series | 32411 |
Number of Images | 5,586,493 |
| Images Size (TB) | 1.6 |
MRI Studies
Trial time points for MRI studies
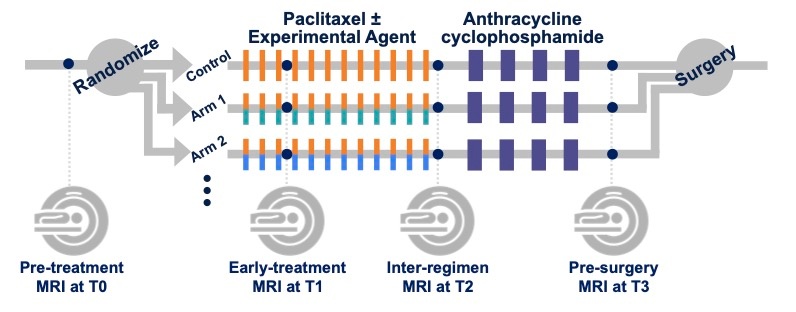
MRI studies were performed at up to four time points in the course of NAC:
- Pre-treatment (T0, optional test/retest visit)
Prior to randomization patients received an MRI study including T2weighted, DWI, and dynamic contrast-enhanced (DCE) acquisitions.
For patients randomized onto a treatment or control arm of I‑SPY 2 subsequent MRI studies with the same required acquisitions were performed:
- Early-treatment (T1, optional test/retest visit)
After 3 weeks of regimen 1 treatment (Paclitaxel with or without an experimental agent). - Mid-treatment (T2)
Between Paclitaxel and Anthracycline (AC) treatment regimens - Post-treatment (T3)
Following 4 cycles AC and prior to surgery.
Figure 1. I-SPY 2 study schema.
Imaging protocol and included original image series
MR imaging was performed on a 1.5 or 3.0 Tesla scanner using a dedicated breast radiofrequency coil. All studies for a given patient were required to be performed on the same scanner configuration (model, field strength and breast coil model). Subjects were imaged in the prone position. The image acquisition protocol included a localization scan and two acquisitions:
- T2weighted sequence
- T1weighted DCE (80sec < phase duration < 100sec with at least 8 minutes continuous post-injection acquisition)
The DWI acquisitions (b=0, 100, 600, 800 s/mm2 OR b=0, 800) were also acquired but are undergoing quality review and are not included in the collection at this time.
Protocol limits on scan parameters for these required acquisitions are given in an accompanying document
All required imaging was performed axially with full bilaterally coverage. For the TCIA collection all original images from the two acquisitions are included. Derived images submitted with the study including reformats, CAD software outputs and MIP images are not part of the collection. For the T2weighted acquisitions multiple series are included if they were generated as part of the standard reconstruction process on the scanner. For example, acquisitions utilizing Dixon techniques to create separate fat and water images will have multiple T2weighted series in the collection.
Derived object series:
Derived objects from the DCE acquisitions are included for the convenience of the user. Some of these are not strictly DICOM compliant and may not be readable by all DICOM software packages. Parametric maps and segmentations are believed to be identical to those utilized in the primary analyses of the I-SPY 2 Trial, but in some cases modifications may have occurred in subsequent processing.
DCE “Package” for each study
- Single-breast ipsilateral cropped DCE images
- As this was done primarily for computational efficiency, low matrix-sized images (<384 voxels per row) were stored and processed without cropping
- All maps and segmentations are based on these cropped images
- Enhancement maps
- PEearly: Percent enhancement at effective post-contrast time 120-150 sec
- PElate: Percent enhancement at effective post-contrast time ~450 sec
- SER: Signal enhancement ratio SER = PEearly / PElate
Note: SER values were scaled by 1000 to allow integer storage. DICOM software utilizing the rescale slope and intercept fields [ DICOM fields (0028, 1052-1054)] should return the original SER values in the range from 0.0 – 3.0. - All maps are stored as DICOM MRI objects
- Functional Tumor Volume (FTV) analysis mask
- Image storing FTV masking processes on a pixel-by-pixel basis. The mask encodes:
- Pre-contrast background thresholding
- Minimum PE thresholding (70% nom.)
- Manually defined rectangular volume of interest (VOI) for enhancing tumor analysis
- Manually defined “OMIT” regions used to exclude non-tumor enhancing regions that encroached on the rectangular VOI
- Stored as DICOM SEG object in a separate series (see Table 1)
- Download Analysis mask files description for further information:
- Image storing FTV masking processes on a pixel-by-pixel basis. The mask encodes:
These DCE derived series can be identified either by Series Description (0008,103e) or by Series Number (0020,0011). See Table 1 for series identification details.
Table 1. Series identification for derived DCE objects
Data type | Series number | Series Description Match |
DCE root directory # | Siemens, GEMS: <root> = 1st original DCE series number Philips: <root> = Original DCE series number / 100 | |
|---|---|---|
SER Map | <root>1000 | ISPY2: VOLSER: uni-lateral cropped: SER 2 |
PE phase <n> Map | <root>100<n> | ISPY2: VOLSER: uni-lateral cropped: PE<n> 2 |
Ipsilateral cropped Original | <root>1800 | ISPY2: VOLSER: uni-lateral cropped: original DCE 2 |
FTV Analysis Mask1 | <root>1900 | ISPY2: VOLSER: uni-lateral cropped: Analysis Mask 2 |
- FTV Analysis Masks are bit-encoded segmentations encoding all masking steps in the I‑SPY 2 primary FTV analysis. See document Analysis mask files description for a full description of these objects.
- DCE images with less than 384 voxels/row were not cropped for the FTV analysis. For these studies the derived DCE objects are all full bilateral images.
DICOM Dictionary for UCSF Private Attributes in Derived Objects:
Calculation parameters and DCE volume analysis results are included in private attributes in some of the derived objects. ACRIN 6698 ISPY2 Shared Private Tag Data Dictionary and ACRIN 6698 ISPY2 DWI and DCE MRI Data Descriptions files are provided for download for researchers wishing to use these parameters and/or results from the primary analyses. Note: the DCE functional tumor volume (FTV) results stored in the SER derived objects are NOT identical to those used in the I‑SPY 2 study due to differences in implementations on different platforms (UCSF in-house software vs. Hologic Inc. AEGIS system). While the differences are generally small and the AEGIS system was validated against the UCSF results in the prior I‑SPY 1 TRIAL, identical results cannot be guaranteed.
Citations & Data Usage Policy
Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution should include references to the following citations:
Data Citation
Li, W., Newitt, D. C., Gibbs, J., Wilmes, L. J., Jones, E. F., Arasu, V. A., Strand, F., Onishi, N., Nguyen, A. A.-T., Kornak, J., Joe, B. N., Price, E. R., Ojeda-Fournier, H., Eghtedari, M., Zamora, K. W., Woodard, S. A., Umphrey, H., Bernreuter, W., Nelson, M., … Hylton, N. M. (2022). I-SPY 2 Breast Dynamic Contrast Enhanced MRI Trial (ISPY2) (Version 1) [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/TCIA.D8Z0-9T85
Data Citation ACRIN-6698
Newitt, D. C., Partridge, S. C., Zhang, Z., Gibbs, J., Chenevert, T., Rosen, M., Bolan, P., Marques, H., Romanoff, J., Cimino, L., Joe, B. N., Umphrey, H., Ojeda-Fournier, H., Dogan, B., Oh, K. Y., Abe, H., Drukteinis, J., Esserman, L. J., & Hylton, N. M. (2021). ACRIN 6698/I-SPY2 Breast DWI [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/TCIA.KK02-6D95
Publication Citation
Li, W., Newitt, D. C., Gibbs, J., Wilmes, L. J., Jones, E. F., Arasu, V. A., Strand, F., Onishi, N., Nguyen, A. A.-T., Kornak, J., Joe, B. N., Price, E. R., Ojeda-Fournier, H., Eghtedari, M., Zamora, K. W., Woodard, S. A., Umphrey, H., Bernreuter, W., Nelson, M., … Hylton, N. M. (2020). Predicting breast cancer response to neoadjuvant treatment using multi-feature MRI: results from the I-SPY 2 TRIAL. In npj Breast Cancer (Vol. 6, Issue 1). Springer Science and Business Media LLC. https://doi.org/10.1038/s41523-020-00203-7
TCIA Citation
Clark, K., Vendt, B., Smith, K., Freymann, J., Kirby, J., Koppel, P., Moore, S., Phillips, S., Maffitt, D., Pringle, M., Tarbox, L., & Prior, F. (2013). The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository. In Journal of Digital Imaging (Vol. 26, Issue 6, pp. 1045–1057). Springer Science and Business Media LLC. https://doi.org/10.1007/s10278-013-9622-7
Other Publications Using This Data
TCIA maintains a list of publications which leverage TCIA data. If you have a publication you'd like to add please contact TCIA's Helpdesk.
Note:
On 7/8/2022 the data submitter signed off on using the CC BY 4.0 instead of the CC BY-NC 4.0.
Version 1 (Current): Updated 2022/05/02
| Data Type | Download all or Query/Filter | License |
|---|---|---|
Images (DICOM, 1.6 TB) 719 patients | Note: When DICOM data are downloaded with the NBIA Data Retriever, the app uses "Series Description" values to construct descriptive directory names. In some series for this Collection, characters that are not allowed in directory names are present. So the workaround is to select the “Classic Directory Name” option, which is located above “Select Directory For Downloaded Files.” (Download requires the NBIA Data Retriever) | |
Images (DICOM, 2.4 TB) I-SPY2 Imaging Cohort 1 dataset, 985 patients | ||
Clinical data (CSV) I-SPY2 Imaging Cohort 1 dataset, 985 patients | ||
Multi-feature MRI NACT Data (.xlsx) “Predicting breast cancer response to neoadjuvant treatment using multi-feature MRI: results from the I-SPY 2 TRIAL” cohort, 384 patients |