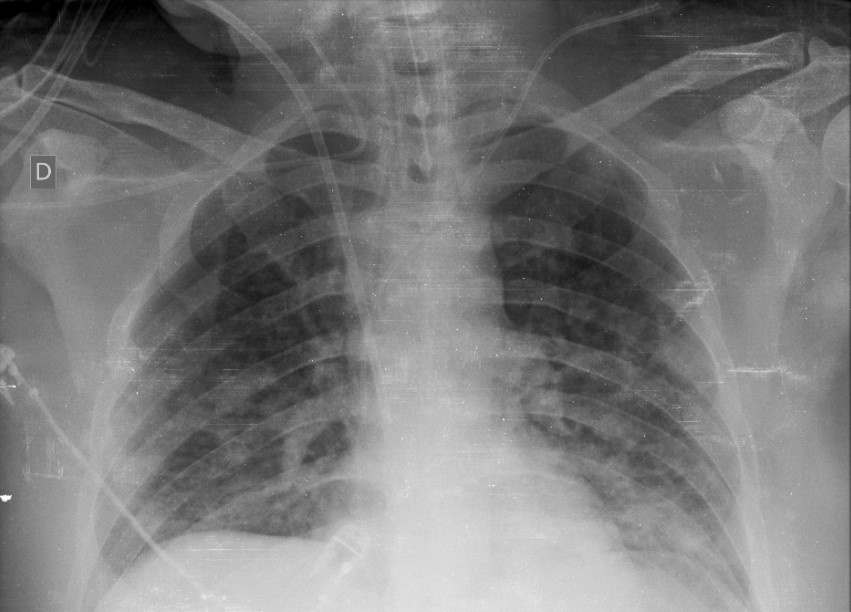
Summary
Background
The COVID-19 pandemic is a global healthcare emergency. Prediction models for COVID-19 imaging are rapidly being developed to support medical decision making in imaging. However, inadequate availability of a diverse annotated dataset has limited the performance and generalizability of existing models.
Purpose
To create the first multi-institutional, multi-national expert annotated COVID-19 imaging dataset made freely available to the machine learning community as a research and educational resource for COVID-19 chest imaging. The Radiological Society of North America (RSNA) assembled the RSNA International COVID-19 Open Radiology Database (RICORD) collection of COVID-related imaging datasets and expert annotations to support research and education. RICORD data will be incorporated in the Medical Imaging and Data Resource Center (MIDRC), a multi-institutional research data repository funded by the National Institute of Biomedical Imaging and Bioengineering of the National Institutes of Health.
Materials and Methods
This dataset was created through a collaboration between the RSNA and Society of Thoracic Radiology (STR). Clinical annotation by thoracic radiology subspecialists was performed for all COVID positive chest radiography (CXR) imaging studies using a labeling schema based upon guidelines for reporting classification of COVID-19 findings in CXRs (see Review of Chest Radiograph Findings of COVID-19 Pneumonia and Suggested Reporting Language, Journal of Thoracic Imaging).
Results
The RSNA International COVID-19 Open Annotated Radiology Database (RICORD) consists of 998 chest x-rays from 361 patients at four international sites annotated with diagnostic labels.
Patient Selection: Patients at least 18 years in age receiving positive diagnosis for COVID-19.
Data Abstract
998 Chest x-ray examinations from 361 patients.
Annotations with labels:
Classification
Typical Appearance
Multifocal bilateral, peripheral opacities, and/or Opacities with rounded morphology
Lower lung-predominant distribution (Required Feature - must be present with either or both of the first two opacity patterns)Indeterminate Appearance
Absence of typical findings AND Unilateral, central or upper lung predominant distribution of airspace disease- Atypical Appearance
Pneumothorax or pleural effusion, Pulmonary Edema, Lobar Consolidation, Solitary lung nodule or mass, Diffuse tiny nodules, Cavity Negative for Pneumonia
No lung opacities
Airspace Disease Grading
Lungs are divided on frontal chest xray into 3 zones per lung (6 zones total). The upper zone extends from the apices to the superior hilum. The mid zone spans between the superior and inferior hilar margins. The lower zone extends from the inferior hilar margins to the costophrenic sulci.Mild - Required if not negative for pneumonia
Opacities in 1-2 lung zonesModerate - Required if not negative for pneumonia
Opacities in 3-4 lung zonesSevere - Required if not negative for pneumonia
Opacities in >4 lung zones
Supporting clinical variables: MRN*, Age, Study Date*, Exam Description, Sex, Study UID*, Image Count, Modality, Testing Result, Specimen Source (* pseudonymous values).
How to use the JSON annotations
More information about how the JSON annotations are organized can be found on https://docs.md.ai/data/json/. Steps 2 & 3 in this example code demonstrate how to to load the JSON into a Dataframe. The JSON file can be downloaded via the data access table below; it is not available via MD.ai. This Jupyter Notebook may also be helpful.
Research Benefits
RICORD is available for non-commercial use (and further enrichment) by the research and education communities which may include development of educational resources for COVID-19, use of RICORD to create AI systems for diagnosis and quantification, benchmarking performance for existing solutions, exploration of distributed/federated learning, further annotation or data augmentation efforts, and evaluation of the examinations for disease entities beyond COVID-19 pneumonia. Deliberate consideration of the detailed annotation schema, demographics, and other included meta-data will be critical when generating cohorts with RICORD, particularly as more public COVID-19 imaging datasets are made available via complementary and parallel efforts. It is important to emphasize that there are limitations to the clinical “ground truth” as the SARS-CoV-2 RT-PCR tests have widely documented limitations and are subject to both false-negative and false-positive results which impact the distribution of the included imaging data, and may have led to an unknown epidemiologic distortion of patients based on the inclusion criteria. These limitations notwithstanding, RICORD has achieved the stated objectives for data complexity, heterogeneity, and high-quality expert annotations as a comprehensive COVID-19 thoracic imaging data resource.
Acknowledgements
We would like to acknowledge the individuals and institutions that have provided data for this collection:
- This dataset was created through a collaboration between the RSNA and Society of Thoracic Radiology (STR). Data in RICORD will be made available through the Medical Imagining Data Resource Center, funded through a contract with the National Institute for Biomedical Imaging and Bioengineering (NIBIB).
Data Access
| Data Type | Download all or Query/Filter | License |
|---|---|---|
Images (DICOM, 11 GB) | (Download requires the NBIA Data Retriever) | |
| Annotations (JSON, 2.34 MB) | ||
| Clinical data (.csv, 88 kB) |
Click the Versions tab for more info about data releases.
Note: The JSON file contains two studies that were subsequently removed: 1.2.826.0.1.3680043.10.474.2925945976491931535320879279573397138 and 1.2.826.0.1.3680043.10.474.2363841256363777216073347285112211043. Please disregard any reference to this data in the JSON file.
Please contact help@cancerimagingarchive.net with any questions regarding usage.
Additional Resources for this Dataset
The NCI Cancer Research Data Commons (CRDC) provides access to additional data and a cloud-based data science infrastructure that connects data sets with analytics tools to allow users to share, integrate, analyze, and visualize cancer research data.
- Imaging Data Commons (IDC) (Imaging Data)
Detailed Description
Image Statistics | |
|---|---|
Modalities | CR, DX |
Number of Patients | 361 |
Number of Studies | 998 |
Number of Series | 1241 |
Number of Images | 1257 |
| Images Size (GB) | 11 |
Citations & Data Usage Policy
Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution should include references to the following citations:
Data Citation
Tsai, E., Simpson, S., Lungren, M.P., Hershman, M., Roshkovan, L., Colak, E., Erickson, B.J., Shih, G., Stein, A.,Kalpathy-Cramer, J., Shen, J.,Hafez, M.A.F., John, S., Rajiah, P., Pogatchnik, B.P., Mongan, J.T., Altinmakas, E., Ranschaert, E., Kitamura, F.C., Topff, L., Moy, L., Kanne, J.P., & Wu, C. (2021). Data from Medical Imaging Data Resource Center (MIDRC) - RSNA International COVID Radiology Database (RICORD) Release 1c - Chest x-ray, Covid+ (MIDRC-RICORD-1C). The Cancer Imaging Archive. DOI: https://doi.org/10.7937/91ah-v663.
Publication Citation
Tsai, E. B., Simpson, S., Lungren, M., Hershman, M., Roshkovan, L., Colak, E., Erickson, B. J., Shih, G., Stein, A., Kalpathy-Cramer, J., Shen, J., Hafez, M., John, S., Rajiah, P., Pogatchnik, B. P., Mongan, J., Altinmakas, E., Ranschaert, E. R., Kitamura, F. C., … Wu, C. C. (2021). The RSNA International COVID-19 Open Annotated Radiology Database (RICORD). Radiology, 203957. DOI: https://doi.org/10.1148/radiol.2021203957
TCIA Citation
Clark K, Vendt B, Smith K, Freymann J, Kirby J, Koppel P, Moore S, Phillips S, Maffitt D, Pringle M, Tarbox L, Prior F. The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository, Journal of Digital Imaging, Volume 26, Number 6, December, 2013, pp 1045-1057. DOI: https://doi.org/10.1007/s10278-013-9622-7
Other Publications Using This Data
TCIA maintains a list of publications which leverage TCIA data. If you have a manuscript you'd like to add please contact the TCIA Helpdesk.
Version 1 (Current): Updated 2021/01/15
| Data Type | Download all or Query/Filter |
|---|---|
Images (DICOM, 11 GB) | (Download requires the NBIA Data Retriever) |
| Annotations (JSON) | |
| Clinical data (.csv) |