Detailed Description | |
|---|
Modalities | MR | Number of Patients | 346 | Number of Studies | 349 | Number of Series | 18,321 | Number of Images | 309,251 | | Images Size (GB) | 15.1 |
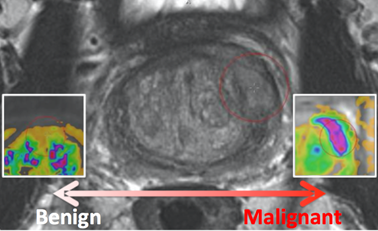
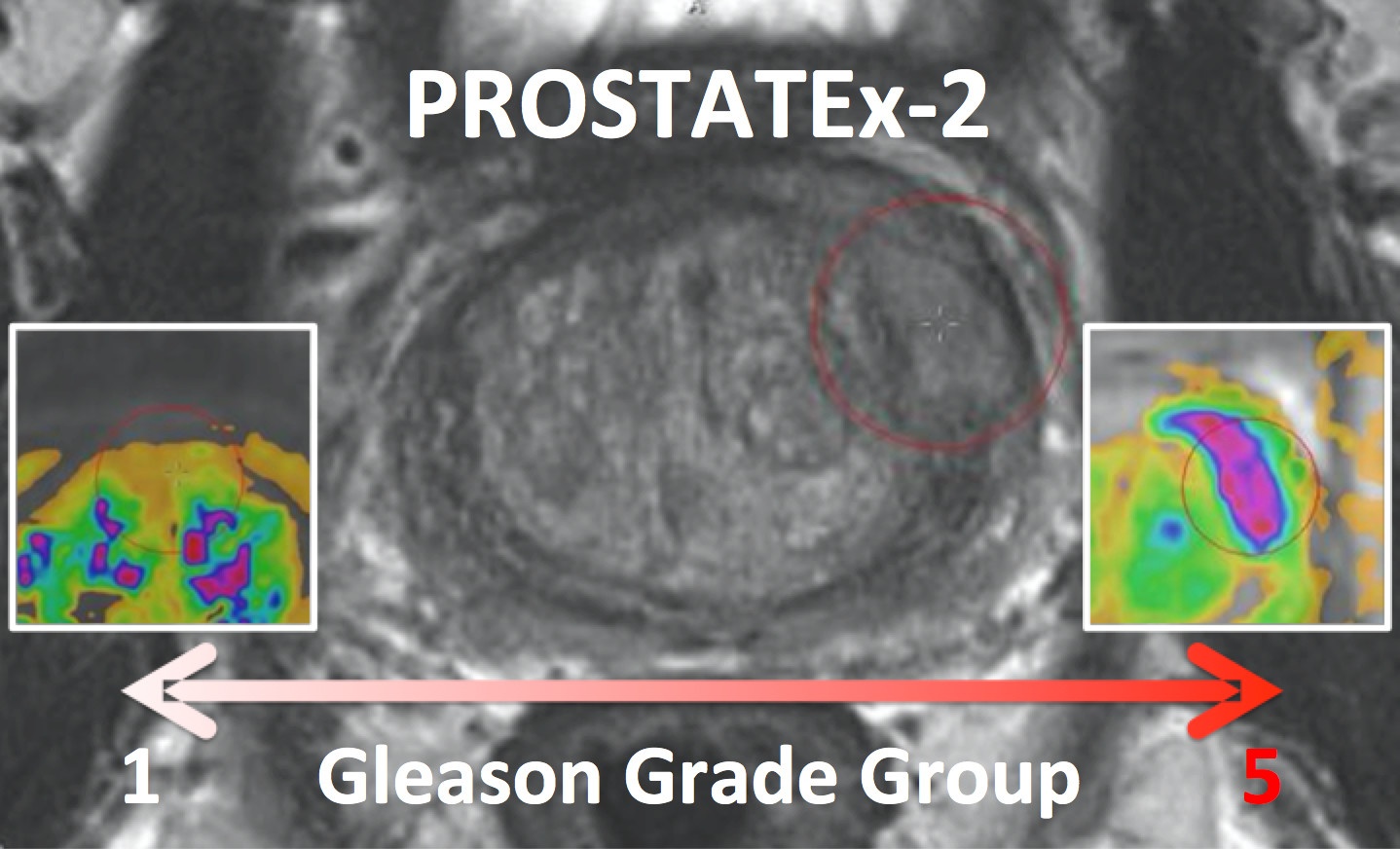
PROSTATEx-2 — SPIE-AAPM-NCI Prostate MR Gleason Grade Group Challenge (May 15, 2017 to August 3, 2017)The American Association of Physicists in Medicine (AAPM), along with the SPIE (the international society for optics and photonics) and the National Cancer Institute (NCI), will conduct a part 2 “Grand Challenge” on the development of quantitative multi-parametric magnetic resonance imaging (MRI) biomarkers for the determination of Gleason Grade Group in prostate cancer. As part of the 2017 AAPM Annual Meeting, the PROSTATEx-2 Challenge will provide a unique opportunity for participants to compare their algorithms with those of others from academia, industry, and government in a structured, direct way using the same data sets. A session at the 2017 AAPM Annual Meeting will focus on the PROSTATEx-2 Challenge; an individual from each of the two top-performing teams will receive a waiver of the meeting registration fee in order to present their methods during this session. Challenge participants are encouraged to submit their work for peer review to the AAPM’s scientific journal Medical Physics. Release date of training set cases with truth: May 15, 2017
Release date of test set cases without truth: June 5, 2017
Submission date for participants’ test set Gleason Grade Group output: June 23, 2017
Challenge results released to participants: June 30, 2017
AAPM Annual Meeting: July 30 – August 3, 2017 (PROSTATEx-2 session: August 1) To participate in PROSATEx-2 you must register at http://spiechallenges.cloudapp.net/competitions/7. PROSTATEx-1 Challenge (November 21, 2016 to February 16, 2017)SPIE, along with the support of the American Association of Physicists in Medicine (AAPM) and the National Cancer Institute (NCI), conducted a “Grand Challenge” on quantitative image analysis methods for the diagnostic classification of clinically significant prostate lesions. As part of the 2017 SPIE Medical Imaging Symposium, the PROSTATEx Challenge provided a unique opportunity for participants to compare their algorithms with those of others from academia, industry, and government in a structured, direct way using the same data sets. For more details, go to http://www.spie.org/PROSTATEx/. - Release date of training set cases with truth: November 21, 2016
- Release date of test set cases without truth: December 12, 2016
- Submission date for participants’ test set classification output: January 16, 2017
- Challenge results released to participants: January 20, 2017
- SPIE Medical Imaging Symposium: February 13-16, 2017
The PROSTATEx Challenge Organizers have provided the following options for download. The significance of these subsets are explained on the the challenge web site: http://spiechallenges.cloudapp.net/competitions/6. | Data Type | Training Cohort (204 subjects) | Test Cohort (140 subjects) |
|---|
Download images (DICOM) | 
|  | Download Ktrans images (.mhd) |  |  | Download lesion information (.zip) |  |  | Download lesion reference thumbnails (.bmp) |  |  |
The images come in two encodings. The acquired MR is provided in DICOM encoding. Additionally Ktrans images are provided. They come in mhd format. Ktrans is a key pharmacokinetic parameter computed from the available Dynamic contrast enhanced T1-weighted series. Each patient has one study with several DICOM images and one Ktrans image. The Ktrans image is encoded in two files ProstateX-[ProxID]-Ktrans.[mhd/zraw], where ProxID is the ProstateX patient identifier. The DICOM images comprise several Series each comprising several Instances. The DICOM files are documented in the ProstateX-Images.csv file. The columns in that file encode the following: ProxID – ProstateX patient identifier. Name – Series Description Studydate – Study Date fid – Finding ID Pos – Scanner Coordinate position of the finding WorldMatrix – Matrix describing image orientation and scaling ijk – image col,row,slice coordinate of finding ImageUID – Image Identifier TopLevel 0 - Series forms one image 1 – A set of Series forms a 4D image (e.g. Dynamic MR) NA – Series form one image, but is part of a Level 1 4D image
SpacingBetweenSlices – Scalar Spacing between slices VoxelSpacing – Vector with x,y,z spacing scalars Dim – Vector with 4D dimensions of the image DCMSerDescr – The original DICOM Series Description DCMSerUID – The DICOM Series UID DCMSerNum – The DICOM Series Number InstanceUIDList – DICOM Instances that make up this series ImageUIDList – TopLevel-NA Images the make up this Toplevel 1 image
For example, to get the ADC image of Patient ProstateX-0123 do the following. After you imported the DICOM files into your environment, go to patient ProstateX-0123 and find the series with ADC in it. In this case it is ‘ep2d_diff_tra_DYNDIST_ADC’. It has SeriesNumber 8. The DICOM images in that series form the ADC image for this challenge. Image slice j at coordinate i,j contains a finding fid. See findings for more details. FindingsThe findings are documented in the ProstateX-Findings.csv table. Documentation for the columns in that table is as follows: pos - Scanner Coordinate position of the finding ClinSig – Identifier available in training set that identifies whether this is a clinically significant finding. Either the biopsy GleasonScore was 7 or higher. Findings with a PIRADS score 2 were not biopsied and are not considered clinically significant. In our center the occurrence of clinically significant cancer in PIRADS 2 lesions is less than 5%.
|