Summary
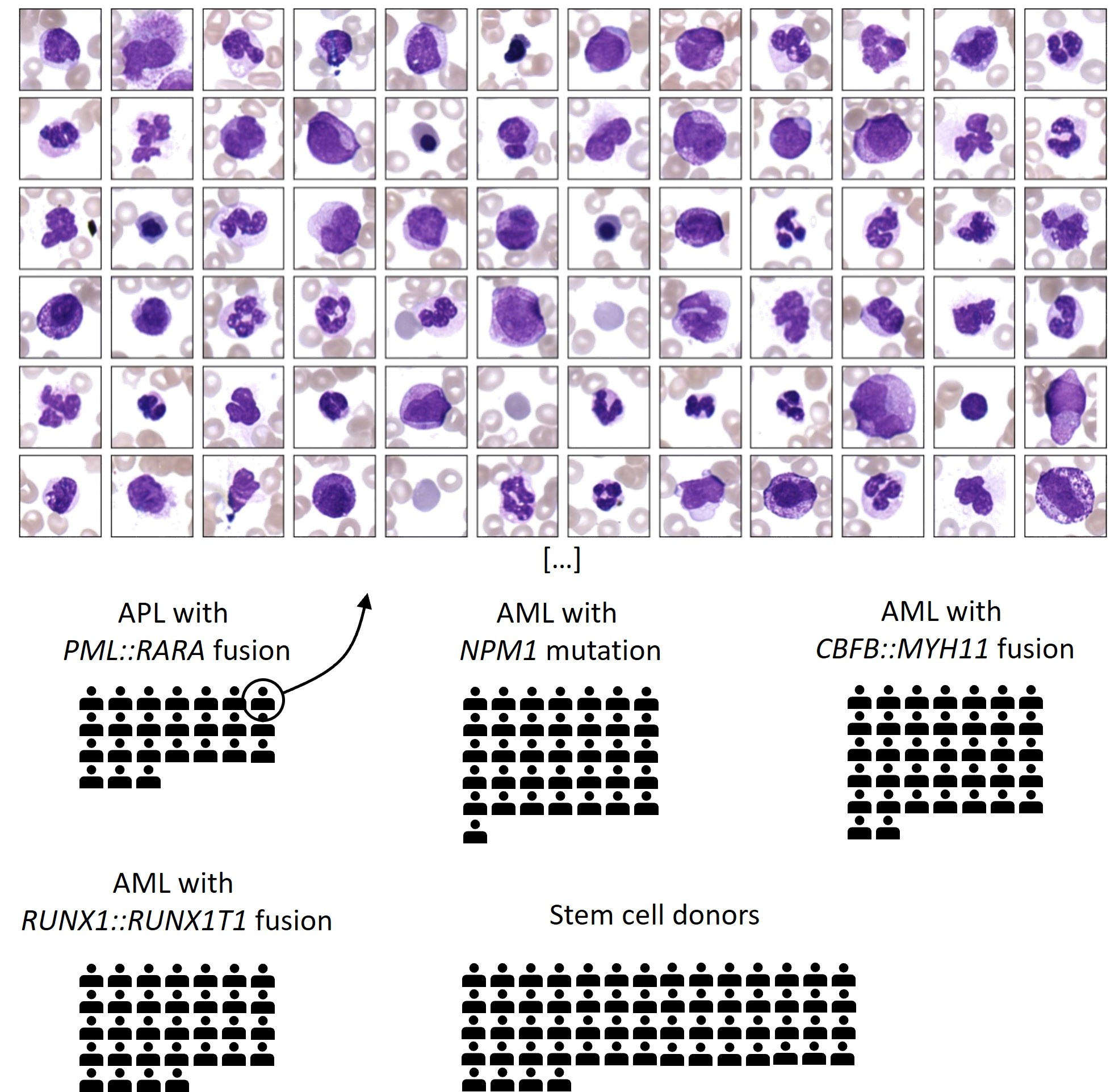
This dataset comprises four prevalent AML subtypes with defining genetic abnormalities and typical morphological features according to the WHO 2022 classification: (i) APL with PML::RARA fusion, (ii) AML with NPM1 mutation, (iii) AML with CBFB::MYH11 fusion (without NPM1 mutation), and (iv) AML with RUNX1::RUNX1T1 fusion, as well as a control group of healthy stem cell donors.A total of 189 peripheral blood smears from the Munich Leukemia Laboratory (MLL) database from the years 2009 to 2020 were digitized. First, all blood smears were scanned with 10x magnification and an overview image was created. Using the Metasystems Metafer platform, cell detection was performed automatically using a segmentation threshold and logarithmic color transformation. Further analysis regarding the quality of the region within the blood smear was performed automatically. Per patient, 99-500 white blood cells were then scanned in 40x magnification via oil immersion microscopy in .TIF format, corresponding to 24,9μm x 24,9μm (144x144 pixels). For this, a CMOS Color Camera from MetaSystems with a resolution of 4096x3000px and a pixel size of 3,45μm x 3,45μm was used. Four pixels were binned into one, leading to a size of 6.9μm x 6.9μm, and a resolution of 6.9μm / 40 (1px = 0,1725μm). Additional information about patient age, sex and blood counts are provided in a separate .csv file.
To our knowledge, this dataset covers the morphological complexity of acute myeloid leukemia in peripheral blood smears in unseen quality and quantity.
Acknowledgements
- All samples were collected, diagnosed and scanned at the Munich Leukemia Laboratory (MLL). Carsten Marr has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (Grant agreement No. 866411). Matthias Hehr acknowledges support from Deutsche José Carreras-Leukämie Stiftung.
Data Access
| Data Type | Download all or Query/Filter | License |
|---|---|---|
| Slide Images (TIFF, 13.3 GB) | (Download requires Aspera) | |
| Clinical metadata (XLS, 12 KB) |
Click the Versions tab for more info about data releases.
Detailed Description
Image Statistics | Pathology Image Statistics |
|---|---|
Modalities | Pathology |
Number of Patients | 189 |
Number of Images | 81,214 |
| Images Size (GB) | 13.3 |
Citations & Data Usage Policy
Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution should include references to the following citations:
Data Citation
Hehr, M., Sadafi, A., Matek, C., Lienemann, P., Pohlkamp, C., Haferlach, T., Spiekermann, K., & Marr, C. (2023). A morphological dataset of white blood cells from patients with four different genetic AML entities and non-malignant controls (AML-Cytomorphology_MLL_Helmholtz) (Version 1) [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/6PPE-4020
Publication Citation
Hehr, M., Sadafi, A., Matek, C., Lienemann, P., Pohlkamp, C., Haferlach, T., Spiekermann, K., & Marr, C. (2023). Explainable AI identifies diagnostic cells of genetic AML subtypes. In H. Mattie (Ed.), PLOS Digital Health (Vol. 2, Issue 3, p. e0000187). Public Library of Science (PLoS). https://doi.org/10.1371/journal.pdig.0000187
TCIA Citation
Clark, K., Vendt, B., Smith, K., Freymann, J., Kirby, J., Koppel, P., Moore, S., Phillips, S., Maffitt, D., Pringle, M., Tarbox, L., & Prior, F. (2013). The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository. In Journal of Digital Imaging (Vol. 26, Issue 6, pp. 1045–1057). Springer Science and Business Media LLC. https://doi.org/10.1007/s10278-013-9622-7
Other Publications Using This Data
TCIA maintains a list of publications which leverage TCIA data. If you have a manuscript you'd like to add please contact the TCIA Helpdesk.